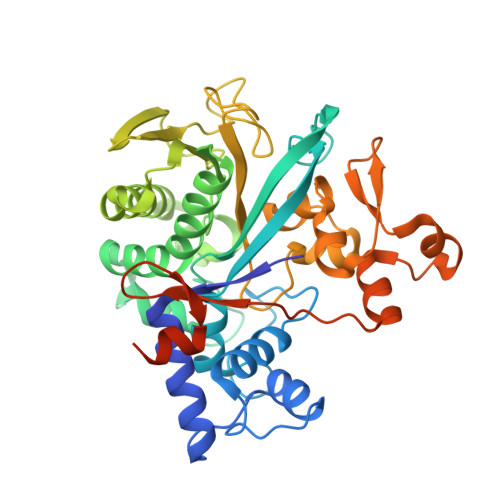
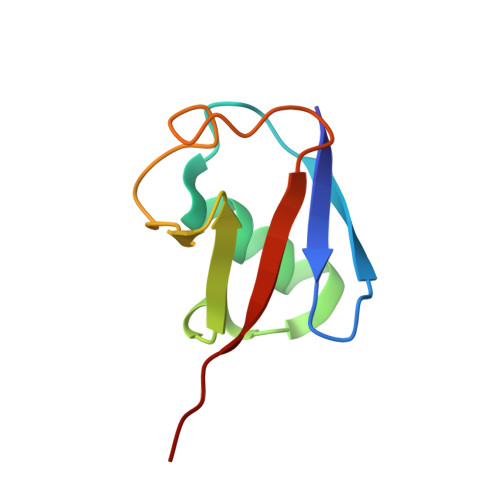
Legionella metaeffector MavL reverses ubiquitin ADP-ribosylation via a conserved arginine-specific macrodomain.
Zhang, Z., Fu, J., Rack, J.G.M., Li, C., Voorneveld, J., Filippov, D.V., Ahel, I., Luo, Z.Q., Das, C.(2024) Nat Commun 15: 2452-2452
- PubMed: 38503748 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-024-46649-2
- Primary Citation Related Structures:
8DMP, 8DMQ, 8DMR, 8DMS, 8DMT, 8DMU - PubMed Abstract:
ADP-ribosylation is a reversible post-translational modification involved in various cellular activities. Removal of ADP-ribosylation requires (ADP-ribosyl)hydrolases, with macrodomain enzymes being a major family in this category. The pathogen Legionella pneumophila mediates atypical ubiquitination of host targets using the SidE effector family in a process that involves ubiquitin ADP-ribosylation on arginine 42 as an obligatory step. Here, we show that the Legionella macrodomain effector MavL regulates this pathway by reversing the arginine ADP-ribosylation, likely to minimize potential detrimental effects caused by the modified ubiquitin. We determine the crystal structure of ADP-ribose-bound MavL, providing structural insights into recognition of the ADP-ribosyl group and catalytic mechanism of its removal. Further analyses reveal DUF4804 as a class of MavL-like macrodomain enzymes whose representative members show unique selectivity for mono-ADP-ribosylated arginine residue in synthetic substrates. We find such enzymes are also present in eukaryotes, as exemplified by two previously uncharacterized (ADP-ribosyl)hydrolases in Drosophila melanogaster. Crystal structures of several proteins in this class provide insights into arginine specificity and a shared mode of ADP-ribose interaction distinct from previously characterized macrodomains. Collectively, our study reveals a new regulatory layer of SidE-catalyzed ubiquitination and expands the current understanding of macrodomain enzymes.
- Department of Chemistry, Purdue University, West Lafayette, IN, 47907, USA.
Organizational Affiliation: