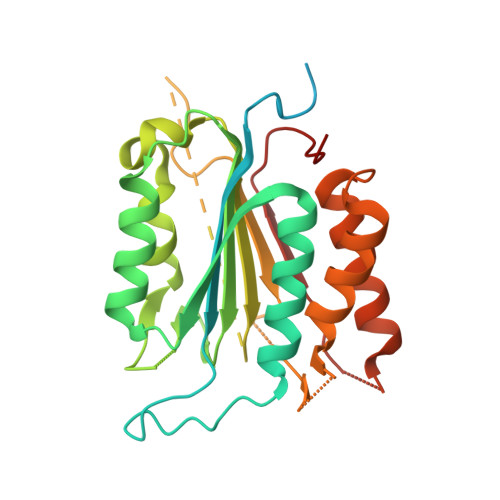
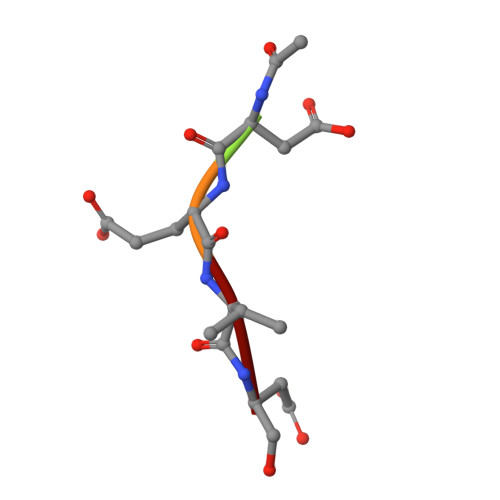
Allosteric Tuning of Caspase-7: Establishing the Nexus of Structure and Catalytic Power.
Hobbs, K.F., Propp, J., Vance, N.R., Kalenkiewicz, A., Witkin, K.R., Ashley Spies, M.(2023) Chemistry 29: e202300872-e202300872
- PubMed: 37005499 Search on PubMed
- DOI: https://doi.org/10.1002/chem.202300872
- Primary Citation Related Structures:
8DGZ, 8DJ3 - PubMed Abstract:
Caspase-7 (C7), a cysteine protease involved in apoptosis, is a valuable drug target for its role in human diseases (e. g., Parkinson's, Alzheimer's, sepsis). The C7 allosteric site has great potential for small-molecule targeting, but numerous drug discovery efforts have identified precious few allosteric inhibitors. Here we present the first selective, drug-like inhibitor of C7 along with several other improved inhibitors based on our previous fragment hit. We also provide a rational basis for the impact of allosteric binding on the C7 catalytic cycle by using an integrated approach including X-ray crystallography, stopped-flow kinetics, and molecular dynamics simulations. Our findings suggest allosteric binding disrupts C7 pre-acylation by neutralization of the catalytic dyad, displacement of substrate from the oxyanion hole, and altered dynamics of substrate binding loops. This work advances drug targeting efforts and bolsters our understanding of allosteric structure-activity relationships (ASARs).
- Biochemistry and Molecular Biology Department, University of Iowa, 51 Newton Road, 4-403 Bowen Science Building, Iowa City, IA, 52242, USA.
Organizational Affiliation: