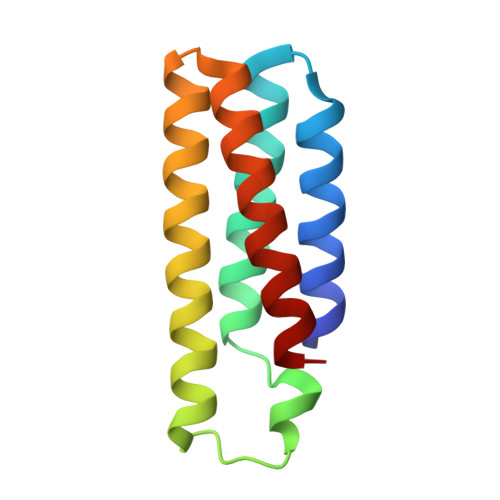
Computationally Guided Redesign of a Heme-free Cytochrome with Native-like Structure and Stability.
Hoffnagle, A.M., Eng, V.H., Markel, U., Tezcan, F.A.(2022) Biochemistry 61: 2063-2072
- PubMed: 36106943 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/acs.biochem.2c00369
- Primary Citation Related Structures:
8DEL, 8DEN - PubMed Abstract:
Metals can play key roles in stabilizing protein structures, but ensuring their proper incorporation is a challenge when a metalloprotein is overexpressed in a non-native cellular environment. Here, we have used computational protein design tools to redesign cytochrome b 562 (cyt b 562 ), which relies on the binding of its heme cofactor to achieve its proper fold, into a stable, heme-free protein. The resulting protein, ApoCyt, features only four mutations and no metal-ligand or covalent bonds, yet displays improved stability over cyt b 562 . Mutagenesis studies and X-ray crystal structures reveal that the increase in stability is due to the computationally prescribed mutations, which stabilize the protein fold through a combination of hydrophobic packing interactions, hydrogen bonds, and cation-π interactions. Upon installation of the relevant mutations, ApoCyt is capable of assembling into previously reported, cytochrome-based trimeric and tetrameric assemblies, demonstrating that ApoCyt retains the structure and assembly properties of cyt b 562 . The successful design of ApoCyt therefore enables further functional diversification of cytochrome-based assemblies and demonstrates that structural metal cofactors can be replaced by a small number of well-designed, non-covalent interactions.
- Department of Chemistry and Biochemistry, University of California, La Jolla, San Diego, California 92093, United States.
Organizational Affiliation: