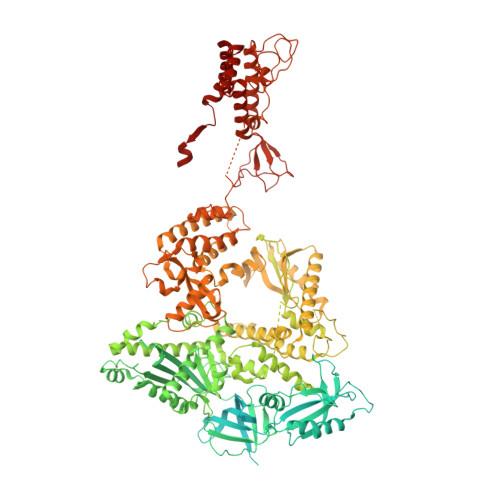
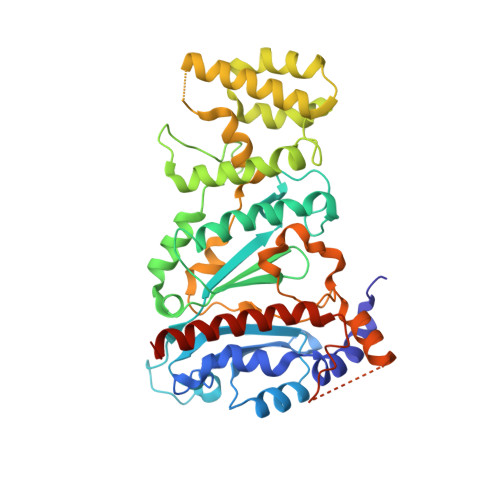
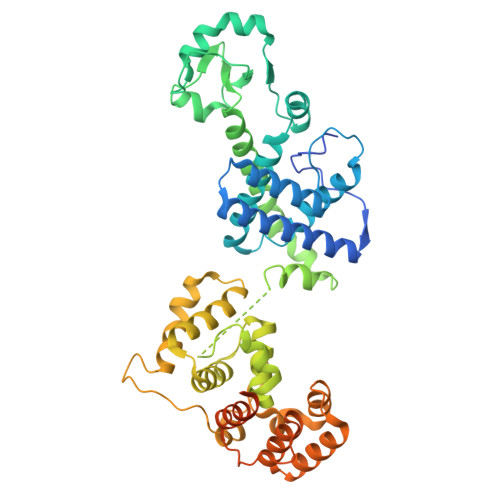
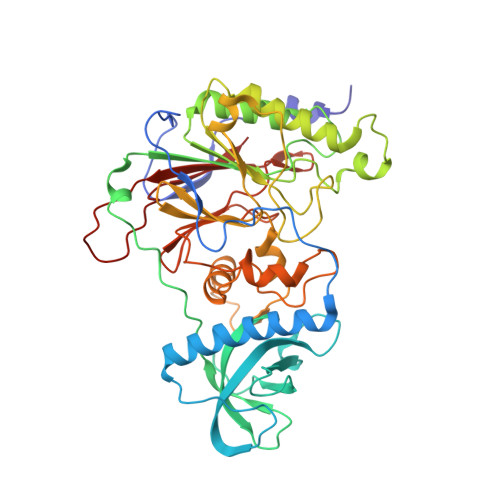
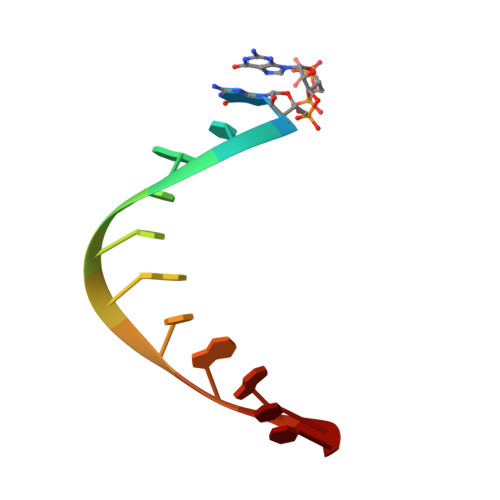
Structures of human primosome elongation complexes.
He, Q., Baranovskiy, A.G., Morstadt, L.M., Lisova, A.E., Babayeva, N.D., Lusk, B.L., Lim, C.J., Tahirov, T.H.(2023) Nat Struct Mol Biol 30: 579-583
- PubMed: 37069376 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41594-023-00971-3
- Primary Citation Related Structures:
8D96, 8D9D - PubMed Abstract:
The synthesis of RNA-DNA primer by primosome requires coordination between primase and DNA polymerase α subunits, which is accompanied by unknown architectural rearrangements of multiple domains. Using cryogenic electron microscopy, we solved a 3.6 Å human primosome structure caught at an early stage of RNA primer elongation with deoxynucleotides. The structure confirms a long-standing role of primase large subunit and reveals new insights into how primosome is limited to synthesizing short RNA-DNA primers.
- Department of Biochemistry, University of Wisconsin-Madison, Madison, WI, USA.
Organizational Affiliation: