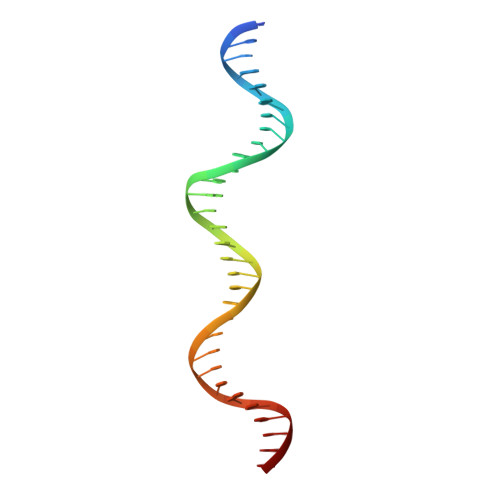
Modular Protein-DNA Cocrystals as Precise, Programmable Assembly Scaffolds.
Orun, A.R., Shields, E.T., Dmytriw, S., Vajapayajula, A., Slaughter, C.K., Snow, C.D.(2023) ACS Nano 17: 13110-13120
- PubMed: 37407546 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/acsnano.2c07282
- Primary Citation Related Structures:
7SDP, 7SOZ, 7U6K, 7U7O, 7UFX, 7UOG, 7UR0, 7UV6, 7UV7, 7UXY, 8D86 - PubMed Abstract:
High-precision nanomaterials to entrap DNA-binding molecules are sought after for applications such as controlled drug delivery and scaffold-assisted structural biology. Here, we engineered protein-DNA cocrystals to serve as scaffolds for DNA-binding molecules. The designed cocrystals, isoreticular cocrystals, contain DNA-binding protein and cognate DNA blocks where the DNA-DNA junctions stack end-to-end. Furthermore, the crystal symmetry allows topology preserving (isoreticular) expansion of the DNA stack without breaking protein-protein contacts, hence providing larger solvent channels for guest diffusion. Experimentally, the resulting designed isoreticular cocrystal adopted an interpenetrating I 222 lattice, a phenomenon previously observed in metal-organic frameworks (MOFs). The interpenetrating lattice crystallized dependably in the same space group despite myriad modifications at the DNA-DNA junctions. Assembly was modular with respect to the DNA inserted for expansion, providing an interchangeable DNA sequence for guest-specified scaffolding. Also, the DNA-DNA junctions were tunable, accommodating varied sticky base overhang lengths and terminal phosphorylation. As a proof of concept, we used the interpenetrating scaffold crystals to separately entrap three distinct guest molecules during crystallization. Isoreticular cocrystal design offers a route to a programmable scaffold for DNA-binding molecules, and the design principles may be applied to existing cocrystals to develop scaffolding materials.
- Department of Chemistry, Colorado State University, 1301 Center Ave., Fort Collins, Colorado 80523, United States.
Organizational Affiliation: