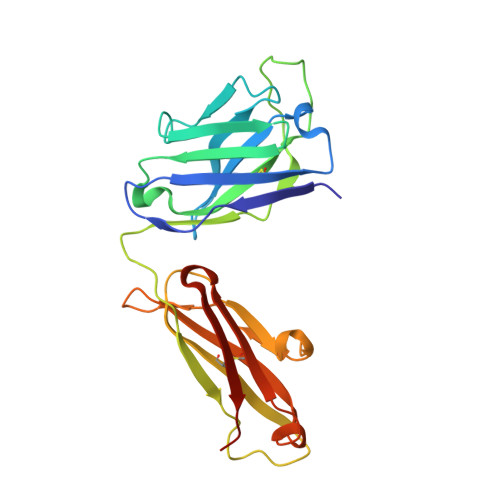
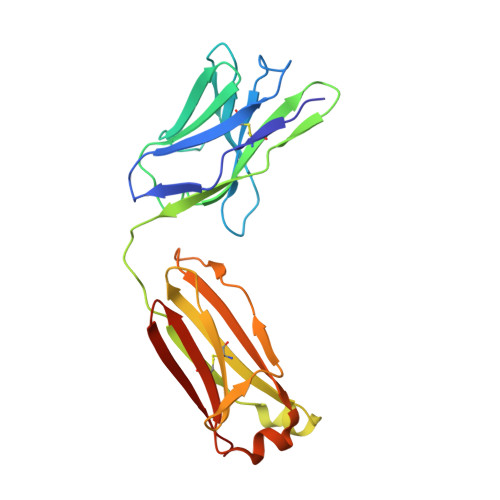
Analysis of two cooperating antibodies unveils immune pressure imposed on HIV Env to elicit a V3-glycan supersite broadly neutralizing antibody lineage.
Finkelstein, M.T., Parker Miller, E., Erdman, M.C., Fera, D.(2022) Front Immunol 13: 962939-962939
- PubMed: 36225920 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.3389/fimmu.2022.962939
- Primary Citation Related Structures:
8D3A - PubMed Abstract:
Elicitation of broadly neutralizing antibodies (bnAbs) is a goal of vaccine design as a strategy for targeting highly divergent strains of HIV-1. Current HIV-1 vaccine design efforts seek to elicit bnAbs by first eliciting their precursors through prime-boost regimens. This requires an understanding of the co-evolution between viruses and antibodies. Towards this goal, we have analyzed two cooperating antibodies, DH475 and DH272, which exerted pressure on the HIV population in an infected donor, called CH848, to evolve in such a way that it became sensitive to the V3-glycan supersite DH270 bnAb lineage. We obtained a 2.90Å crystal structure of DH475 in complex with the Man 9 glycan and a negative stain EM model of DH272 in complex with the HIV-1 spike trimer, Env. Coupled with additional modeling studies and biochemical data, our studies reveal that DH475 contacts a V3- and V4-glycan dependent epitope accessible on an open or shed Env and that DH272 makes critical contacts with the V1V2 and V3 loops on HIV-1 Env. Using these data, we suggest a prime-boost regimen that may facilitate the initiation of DH270-like bnAb precursors.
- Department of Chemistry and Biochemistry, Swarthmore College, Swarthmore, PA, United States.
Organizational Affiliation: