TBD
Song, X., Butler, J., Li, C., Zhang, K., Zhang, D., Hao, Y.To be published.
Experimental Data Snapshot
Starting Models: experimental
View more details
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Ubiquitin carboxyl-terminal hydrolase 30 | 349 | Homo sapiens | Mutation(s): 3 Gene Names: USP30 EC: 3.4.19.12 | 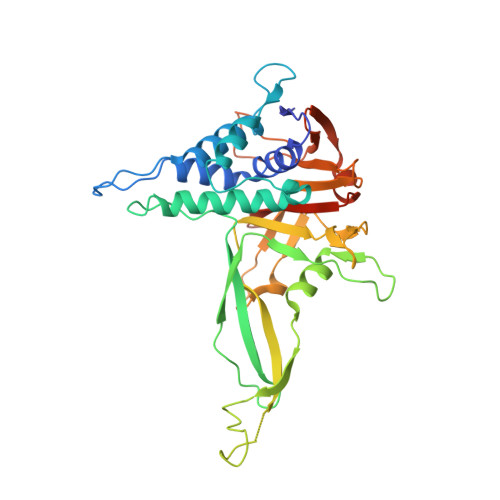 | |
UniProt & NIH Common Fund Data Resources | |||||
PHAROS: Q70CQ3 GTEx: ENSG00000135093 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | Q70CQ3 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| mouse anti-huUSP30 Fab heavy chain | B [auth H] | 222 | Mus musculus | Mutation(s): 0 | 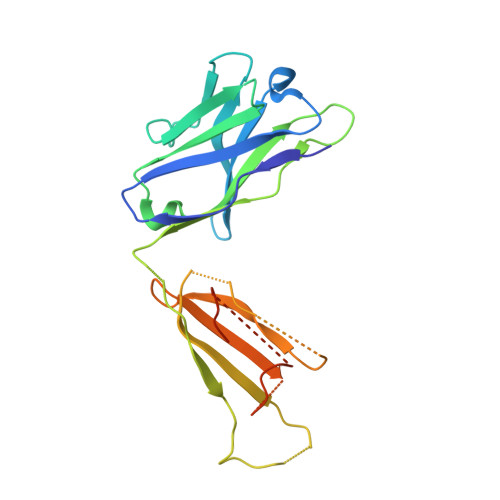 |
Entity ID: 3 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| mouse anti-huUSP30 Fab light chain | C [auth L] | 214 | Mus musculus | Mutation(s): 0 | 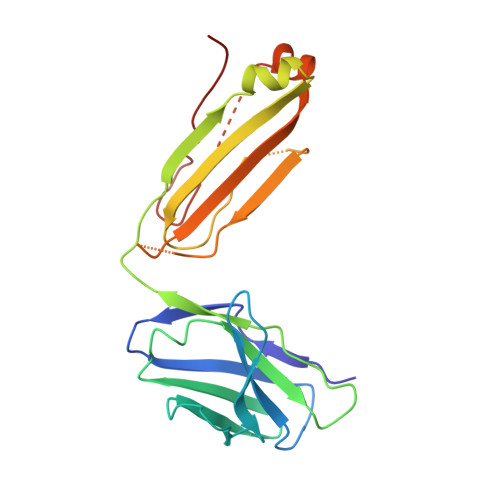 |
| Ligands 3 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| PXW (Subject of Investigation/LOI) Download:Ideal Coordinates CCD File | D [auth A] | (1R,2R,4S,7E)-7-[amino(sulfanyl)methylidene]-2-{[(1P)-3-chloro-3'-(1-cyanocyclopropyl)[1,1'-biphenyl]-4-carbonyl]amino}-7-azabicyclo[2.2.1]heptan-7-ium C24 H24 Cl N4 O S QYJCMFNKFUMQPQ-IOMROCGXSA-O |  | ||
| ZN Download:Ideal Coordinates CCD File | E [auth A] | ZINC ION Zn PTFCDOFLOPIGGS-UHFFFAOYSA-N |  | ||
| EDO Download:Ideal Coordinates CCD File | F [auth A] | 1,2-ETHANEDIOL C2 H6 O2 LYCAIKOWRPUZTN-UHFFFAOYSA-N |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 41.879 | α = 90 |
| b = 71.721 | β = 93.98 |
| c = 148.63 | γ = 90 |
| Software Name | Purpose |
|---|---|
| XDS | data reduction |
| XDS | data scaling |
| PHASER | phasing |
| PHENIX | refinement |
| Funding Organization | Location | Grant Number |
|---|---|---|
| Other private | Amgen Inc | |
| Other private | Carmot Therapeutics Inc |