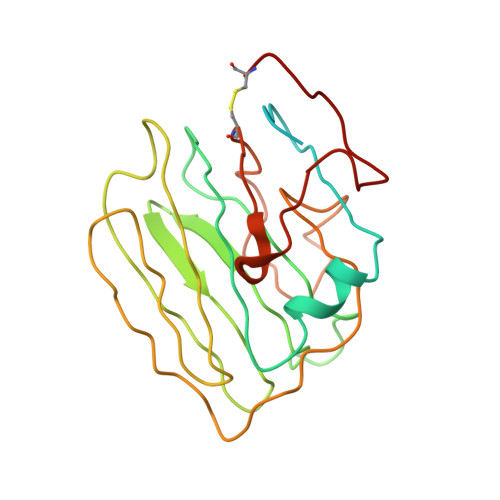
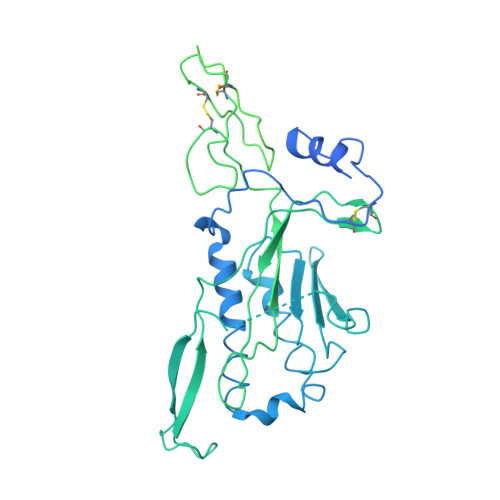
Structure of the photoreceptor synaptic assembly of the extracellular matrix protein pikachurin with the orphan receptor GPR179.
Patil, D.N., Pantalone, S., Cao, Y., Laboute, T., Novick, S.J., Singh, S., Savino, S., Faravelli, S., Magnani, F., Griffin, P.R., Singh, A.K., Forneris, F., Martemyanov, K.A.(2023) Sci Signal 16: eadd9539-eadd9539
- PubMed: 37490546 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1126/scisignal.add9539
- Primary Citation Related Structures:
7ZC9, 7ZCB, 8D1B - PubMed Abstract:
Precise synapse formation is essential for normal functioning of the nervous system. Retinal photoreceptors establish selective contacts with bipolar cells, aligning the neurotransmitter release apparatus with postsynaptic signaling cascades. This involves transsynaptic assembly between the dystroglycan-dystrophin complex on the photoreceptor and the orphan receptor GPR179 on the bipolar cell, which is mediated by the extracellular matrix protein pikachurin (also known as EGFLAM). This complex plays a critical role in the synaptic organization of photoreceptors and signal transmission, and mutations affecting its components cause blinding disorders in humans. Here, we investigated the structural organization and molecular mechanisms by which pikachurin orchestrates transsynaptic assembly and solved structures of the human pikachurin domains by x-ray crystallography and of the GPR179-pikachurin complex by single-particle, cryo-electron microscopy. The structures reveal molecular recognition principles of pikachurin by the Cache domains of GPR179 and show how the interaction is involved in the transsynaptic alignment of the signaling machinery. Together, these data provide a structural basis for understanding the synaptic organization of photoreceptors and ocular pathology.
- Department of Neuroscience, Herbert Wertheim UF Scripps Institute for Biomedical Innovation and Technology, University of Florida, Jupiter, FL 33458, USA.
Organizational Affiliation: