Atropo/Tropo Flexibility: A Tool for Design and Synthesis of Self-Adaptable Inhibitors of Carbonic Anhydrases and Their Antiproliferative Effect.
Ivanova, J.N., Nocentini, A., Ta Rs, K., Leita Ns, J.N., Dvinskis, E., Kazaks, A., Domraceva, I., Supuran, C.T., Zalubovskis, R.(2023) J Med Chem 66: 5703-5718
- PubMed: 37022308 Search on PubMed
- DOI: https://doi.org/10.1021/acs.jmedchem.3c00007
- Primary Citation Related Structures:
8CO0, 8CO3 - PubMed Abstract:
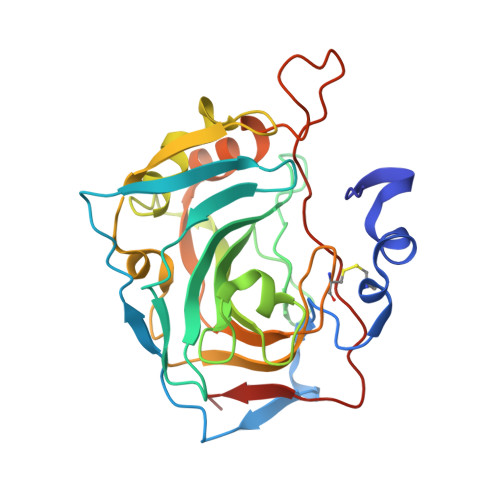
Here, we report for the first time a series of sulfonamide derivatives with scaffolds bearing flexible moieties, namely, rotamers or tropoisomers capable of adapting their geometry in the active center of enzymes thus being effective and selective carbonic anhydrase (CAs, EC 4.2.1.1) enzyme inhibitors. All compounds exhibited effective in vitro inhibition activity toward the main hCA isoforms related to cancer (i.e., hCA II, hCA IX, and hCA XII with K I values in the low nanomolar range). Three selected compounds showed a great cytotoxic effect on cancer cell lines ex vivo. X-ray crystallographic experiments assessed the binding modes of compound 35 with active centers of hCA IX and hCA XII.
- Latvian Institute of Organic Synthesis, Aizkraukles 21, LV-1006 Riga, Latvia.
Organizational Affiliation: