Lessons on protein structure from interleukin-4: All disulfides are not created equal.
Vaz, D.C., Rodrigues, J.R., Loureiro-Ferreira, N., Muller, T.D., Sebald, W., Redfield, C., Brito, R.M.M.(2024) Proteins 92: 219-235
- PubMed: 37814578 Search on PubMed
- DOI: https://doi.org/10.1002/prot.26611
- Primary Citation Related Structures:
8A4F, 8CGF, 8CH7 - PubMed Abstract:
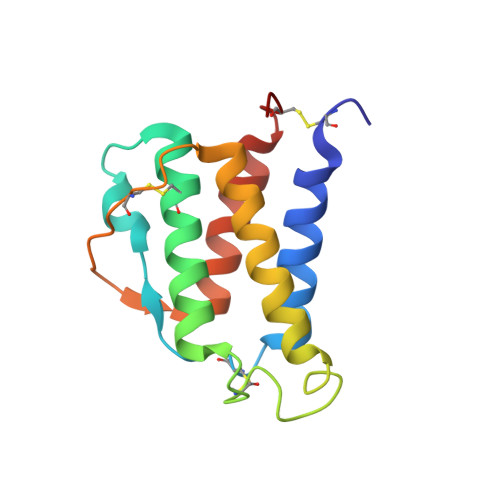
Interleukin-4 (IL-4) is a hematopoietic cytokine composed by a four-helix bundle stabilized by an antiparallel beta-sheet and three disulfide bonds: Cys3-Cys127, Cys24-Cys65, and Cys46-Cys99. IL-4 is involved in several immune responses associated to infection, allergy, autoimmunity, and cancer. Besides its physiological relevance, IL-4 is often used as a "model" for protein design and engineering. Hence, to understand the role of each disulfide in the structure and dynamics of IL-4, we carried out several spectroscopic analyses (circular dichroism [CD], fluorescence, nuclear magnetic resonance [NMR]), and molecular dynamics (MD) simulations on wild-type IL-4 and four IL-4 disulfide mutants. All disulfide mutants showed loss of structure, altered interhelical angles, and looser core packings, showing that all disulfides are relevant for maintaining the overall fold and stability of the four-helix bundle motif, even at very low pH. In the absence of the disulfide connecting both protein termini Cys3-Cys127, C3T-IL4 showed a less packed protein core, loss of secondary structure (~9%) and fast motions on the sub-nanosecond time scale (lower S 2 order parameters and larger τ c correlation time), especially at the two protein termini, loops, beginning of helix A and end of helix D. In the absence of Cys24-Cys65, C24T-IL4 presented shorter alpha-helices (14% loss in helical content), altered interhelical angles, less propensity to form the small anti-parallel beta-sheet and increased dynamics. Simultaneously deprived of two disulfides (Cys3-Cys127 and Cys24-Cys65), IL-4 formed a partially folded "molten globule" with high 8-anilino-1-naphtalenesulphonic acid-binding affinity and considerable loss of secondary structure (~50%decrease), as shown by the far UV-CD, NMR, and MD data.
- School of Health Sciences, Polytechnic of Leiria, Leiria, Portugal.
Organizational Affiliation: