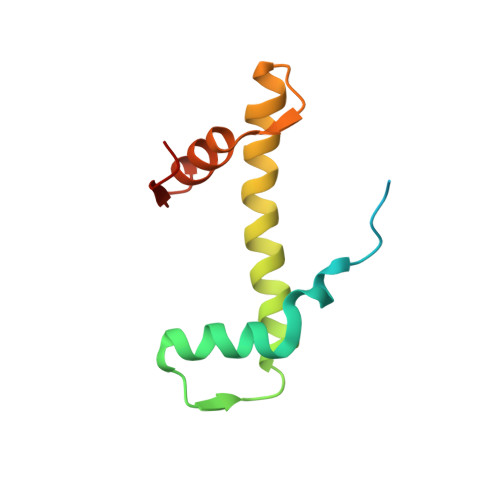
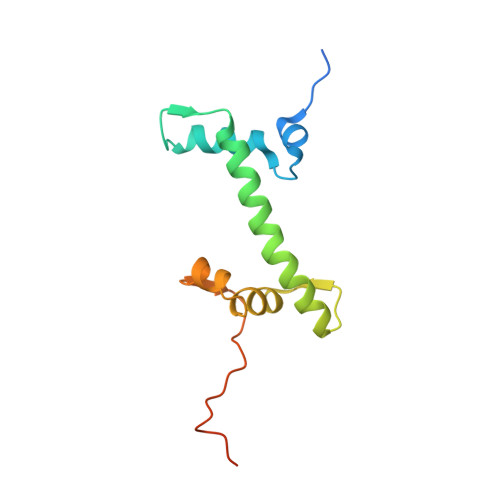
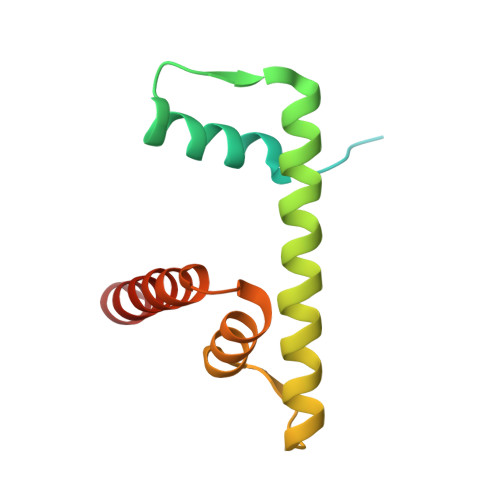
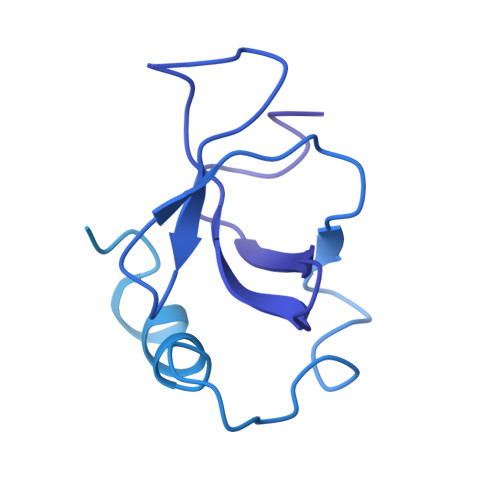
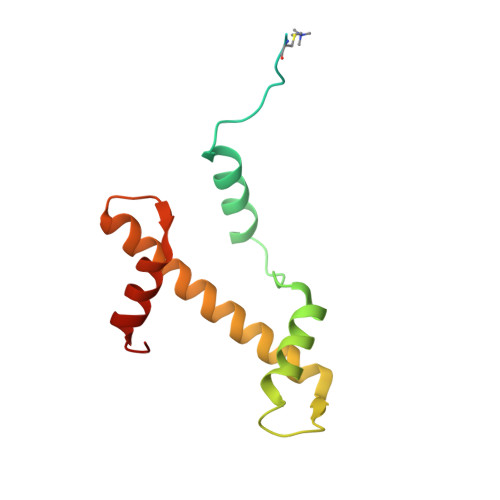
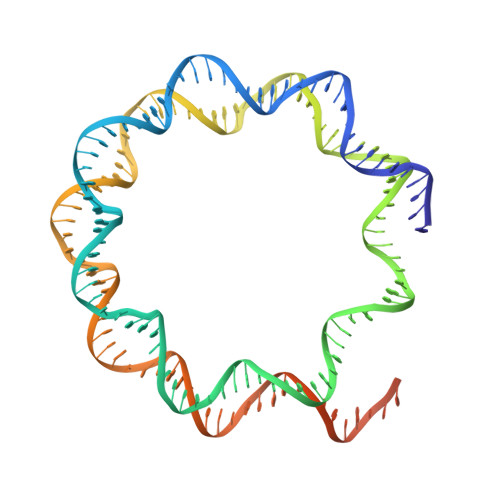
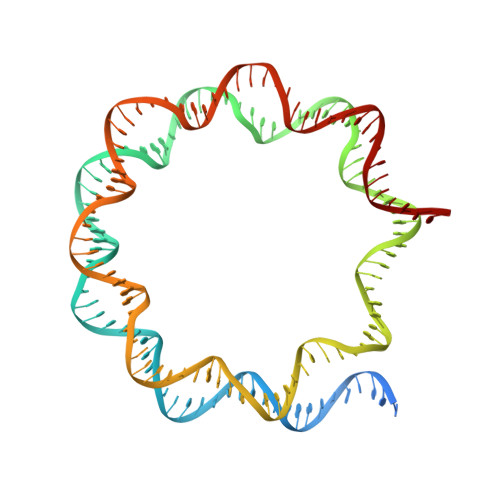
Multivalency of nucleosome recognition by LEDGF.
Koutna, E., Lux, V., Kouba, T., Skerlova, J., Novacek, J., Srb, P., Hexnerova, R., Svachova, H., Kukacka, Z., Novak, P., Fabry, M., Poepsel, S., Veverka, V.(2023) Nucleic Acids Res 51: 10011-10025
- PubMed: 37615563 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gkad674
- Primary Citation Related Structures:
8CBN, 8CBQ, 8PC5, 8PC6, 8PEO, 8PEP - PubMed Abstract:
Eukaryotic transcription is dependent on specific histone modifications. Their recognition by chromatin readers triggers complex processes relying on the coordinated association of transcription regulatory factors. Although various modification states of a particular histone residue often lead to differential outcomes, it is not entirely clear how they are discriminated. Moreover, the contribution of intrinsically disordered regions outside of the specialized reader domains to nucleosome binding remains unexplored. Here, we report the structures of a PWWP domain from transcriptional coactivator LEDGF in complex with the H3K36 di- and trimethylated nucleosome, indicating that both methylation marks are recognized by PWWP in a highly conserved manner. We identify a unique secondary interaction site for the PWWP domain at the interface between the acidic patch and nucleosomal DNA that might contribute to an H3K36-methylation independent role of LEDGF. We reveal DNA interacting motifs in the intrinsically disordered region of LEDGF that discriminate between the intra- or extranucleosomal DNA but remain dynamic in the context of dinucleosomes. The interplay between the LEDGF H3K36-methylation reader and protein binding module mediated by multivalent interactions of the intrinsically disordered linker with chromatin might help direct the elongation machinery to the vicinity of RNA polymerase II, thereby facilitating productive elongation.
- Institute of Organic Chemistry and Biochemistry of the Czech Academy of Sciences, Prague 160 00, Czech Republic.
Organizational Affiliation: