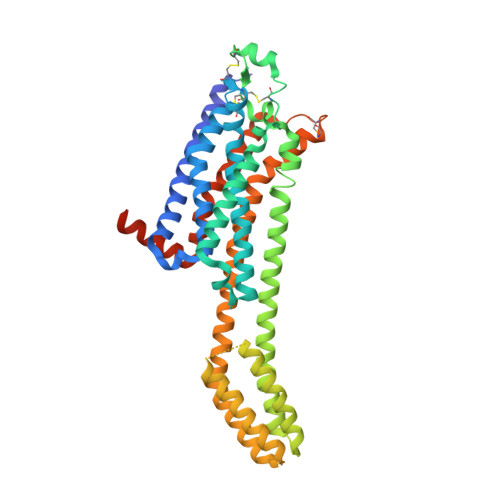
Crystal structure of adenosine A 2A receptor in complex with clinical candidate Etrumadenant reveals unprecedented antagonist interaction.
Claff, T., Schlegel, J.G., Voss, J.H., Vaassen, V.J., Weisse, R.H., Cheng, R.K.Y., Markovic-Mueller, S., Bucher, D., Strater, N., Muller, C.E.(2023) Commun Chem 6: 106-106
- PubMed: 37264098 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s42004-023-00894-6
- Primary Citation Related Structures:
8C9W, 8CIC - PubMed Abstract:
The G s protein-coupled adenosine A 2A receptor (A 2A AR) represents an emerging drug target for cancer immunotherapy. The clinical candidate Etrumadenant was developed as an A 2A AR antagonist with ancillary blockade of the A 2B AR subtype. It constitutes a unique chemotype featuring a poly-substituted 2-amino-4-phenyl-6-triazolylpyrimidine core structure. Herein, we report two crystal structures of the A 2A AR in complex with Etrumadenant, obtained with differently thermostabilized A 2A AR constructs. This led to the discovery of an unprecedented interaction, a hydrogen bond of T88 3.36 with the cyano group of Etrumadenant. T88 3.36 is mutated in most A 2A AR constructs used for crystallization, which has prevented the discovery of its interactions. In-vitro characterization of Etrumadenant indicated low selectivity versus the A 1 AR subtype, which can be rationalized by the structural data. These results will facilitate the future design of AR antagonists with desired selectivity. Moreover, they highlight the advantages of the employed A 2A AR crystallization construct that is devoid of ligand binding site mutations.
- PharmaCenter Bonn & Pharmaceutical Institute, Department of Pharmaceutical & Medicinal Chemistry, University of Bonn, An der Immenburg 4, 53113, Bonn, Germany. tobias.claff@uni-bonn.de.
Organizational Affiliation: