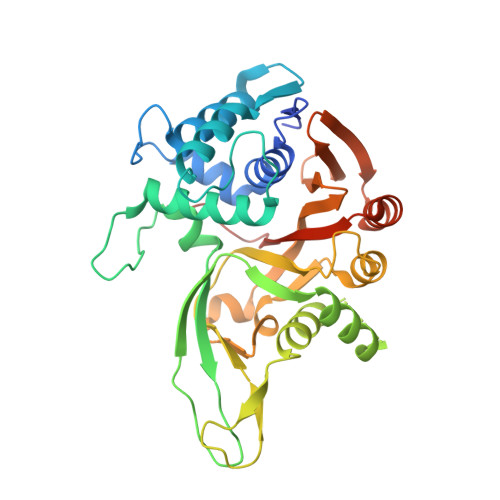
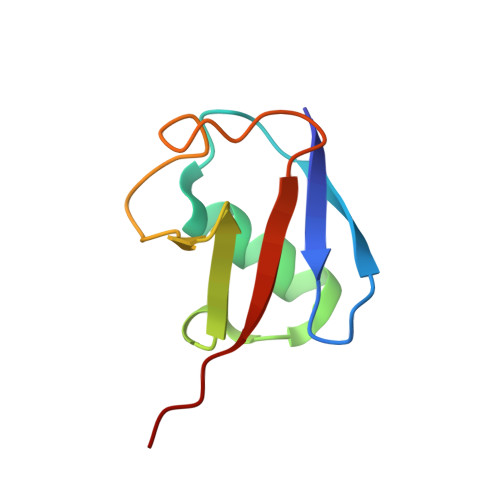
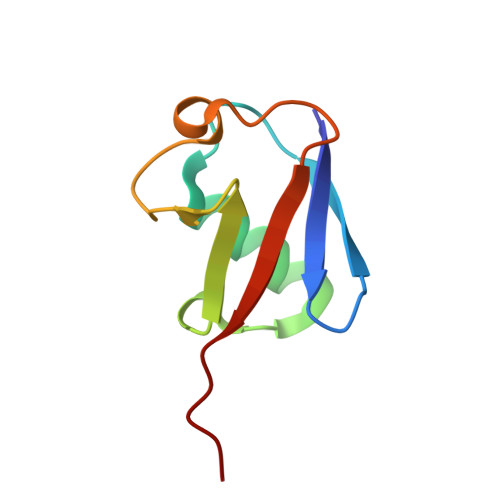
Discovery and mechanism of K63-linkage-directed deubiquitinase activity in USP53.
Wendrich, K., Gallant, K., Recknagel, S., Petroulia, S., Kazi, N.H., Hane, J.A., Fuhrer, S., Bezstarosti, K., O'Dea, R., Demmers, J., Gersch, M.(2024) Nat Chem Biol
- PubMed: 39587316 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41589-024-01777-0
- Primary Citation Related Structures:
8C61 - PubMed Abstract:
Ubiquitin-specific proteases (USPs) represent the largest class of human deubiquitinases (DUBs) and comprise its phylogenetically most distant members USP53 and USP54, which are annotated as catalytically inactive pseudoenzymes. Conspicuously, mutations within the USP domain of USP53 cause progressive familial intrahepatic cholestasis. Here, we report the discovery that USP53 and USP54 are active DUBs with high specificity for K63-linked polyubiquitin. We demonstrate how USP53 mutations abrogate catalytic activity, implicating loss of DUB activity in USP53-mediated pathology. Depletion of USP53 increases K63-linked ubiquitination of tricellular junction components. Assays with substrate-bound polyubiquitin reveal that USP54 cleaves within K63-linked chains, whereas USP53 can en bloc deubiquitinate substrate proteins in a K63-linkage-dependent manner. Biochemical and structural analyses uncover underlying K63-specific S2 ubiquitin-binding sites within their catalytic domains. Collectively, our work revises the annotation of USP53 and USP54, provides reagents and a mechanistic framework to investigate K63-linked polyubiquitin decoding and establishes K63-linkage-directed deubiquitination as a new DUB activity.
- Chemical Genomics Centre, Max Planck Institute of Molecular Physiology, Dortmund, Germany.
Organizational Affiliation: