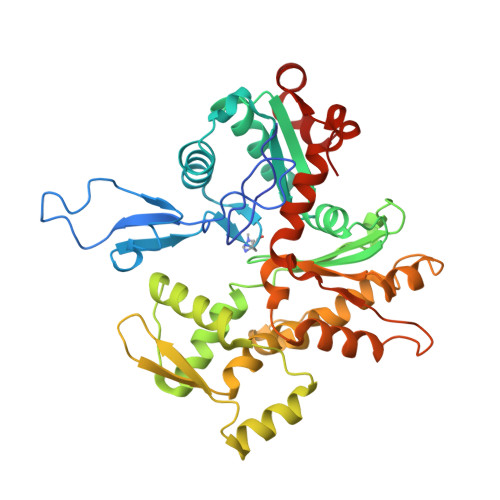
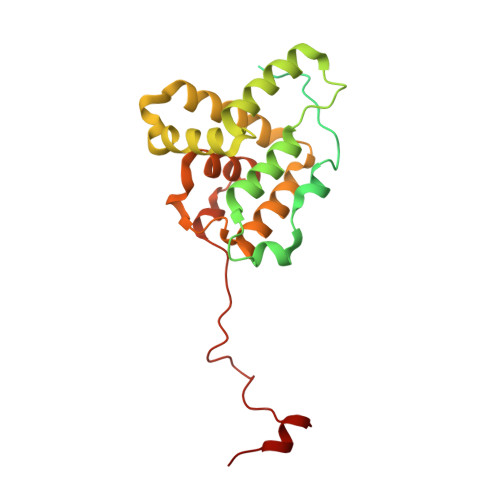
Structural basis for subversion of host cell actin cytoskeleton during Salmonella infection.
Yuan, B., Scholz, J., Wald, J., Thuenauer, R., Hennell James, R., Ellenberg, I., Windhorst, S., Faix, J., Marlovits, T.C.(2023) Sci Adv 9: eadj5777-eadj5777
- PubMed: 38064550 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1126/sciadv.adj5777
- Primary Citation Related Structures:
8C4C, 8C4E - PubMed Abstract:
Secreted bacterial type III secretion system (T3SS) proteins are essential for successful infection by many human pathogens. Both T3SS translocator SipC and effector SipA are critical for Salmonella infection by subversion of the host cell cytoskeleton, but the precise molecular interplay between them remains unknown. Here, using cryo-electron microscopy, we show that SipA binds along the F-actin grooves with a unique binding pattern. SipA stabilizes F-actin through charged interface residues and appears to prevent inorganic phosphate release through closure of the "back door" of adenosine 5'-triphosphate pocket. We also show that SipC enhances the binding of SipA to F-actin, thus demonstrating that a sequential presence of T3SS proteins in host cells is associated with a sequence of infection events-starting with actin nucleation, filament growth, and stabilization. Together, our data explain the coordinated interplay of a precisely tuned and highly effective mechanism during Salmonella infection and provide a blueprint for interfering with Salmonella effectors acting on actin.
- University Medical Center Hamburg-Eppendorf (UKE), Institute of Structural and Systems Biology, Hamburg, Germany.
Organizational Affiliation: