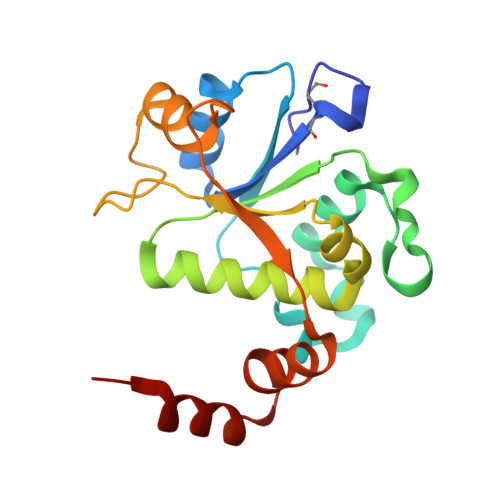
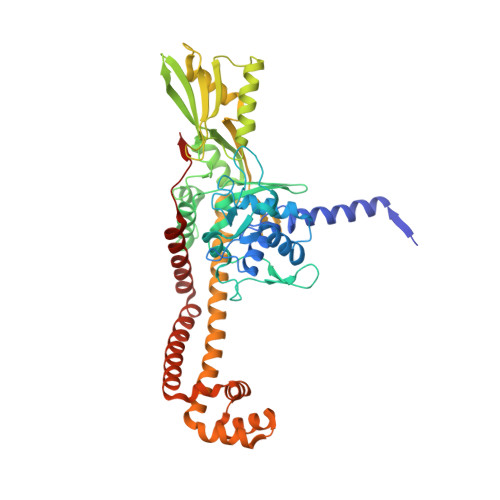
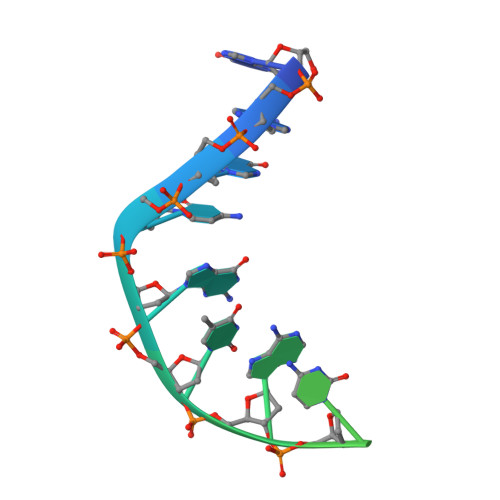
A 2.8 angstrom Structure of Zoliflodacin in a DNA Cleavage Complex with Staphylococcus aureus DNA Gyrase.
Morgan, H., Lipka-Lloyd, M., Warren, A.J., Hughes, N., Holmes, J., Burton, N.P., Mahenthiralingam, E., Bax, B.D.(2023) Int J Mol Sci 24
- PubMed: 36675148 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.3390/ijms24021634
- Primary Citation Related Structures:
8BP2 - PubMed Abstract:
Since 2000, some thirteen quinolones and fluoroquinolones have been developed and have come to market. The quinolones, one of the most successful classes of antibacterial drugs, stabilize DNA cleavage complexes with DNA gyrase and topoisomerase IV (topo IV), the two bacterial type IIA topoisomerases. The dual targeting of gyrase and topo IV helps decrease the likelihood of resistance developing. Here, we report on a 2.8 Å X-ray crystal structure, which shows that zoliflodacin, a spiropyrimidinetrione antibiotic, binds in the same DNA cleavage site(s) as quinolones, sterically blocking DNA religation. The structure shows that zoliflodacin interacts with highly conserved residues on GyrB (and does not use the quinolone water-metal ion bridge to GyrA), suggesting it may be more difficult for bacteria to develop target mediated resistance. We show that zoliflodacin has an MIC of 4 µg/mL against Acinetobacter baumannii ( A. baumannii ), an improvement of four-fold over its progenitor QPT-1. The current phase III clinical trial of zoliflodacin for gonorrhea is due to be read out in 2023. Zoliflodacin, together with the unrelated novel bacterial topoisomerase inhibitor gepotidacin, is likely to become the first entirely novel chemical entities approved against Gram-negative bacteria in the 21st century. Zoliflodacin may also become the progenitor of a new safer class of antibacterial drugs against other problematic Gram-negative bacteria.
- Medicines Discovery Institute, Cardiff University, Cardiff CF10 3AT, UK.
Organizational Affiliation: