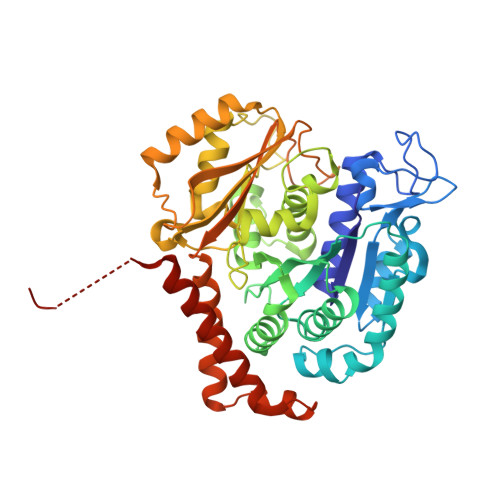
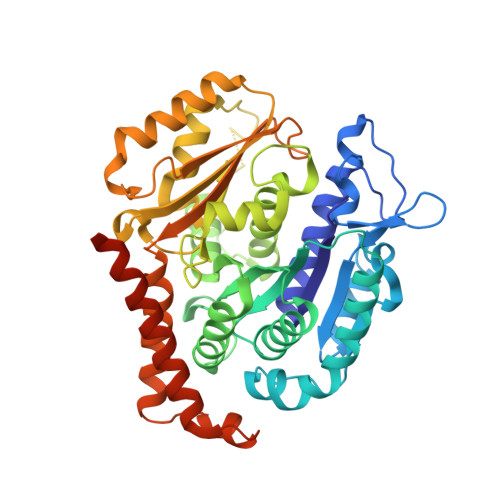
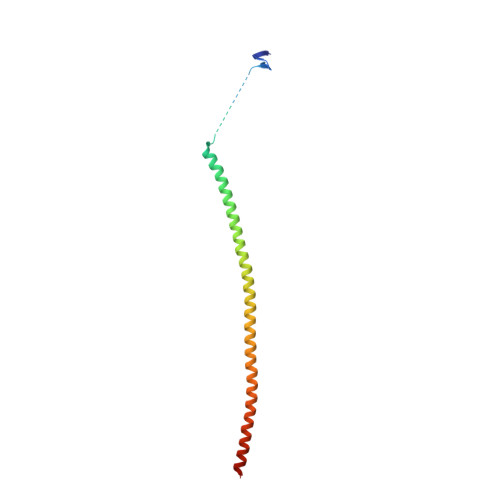
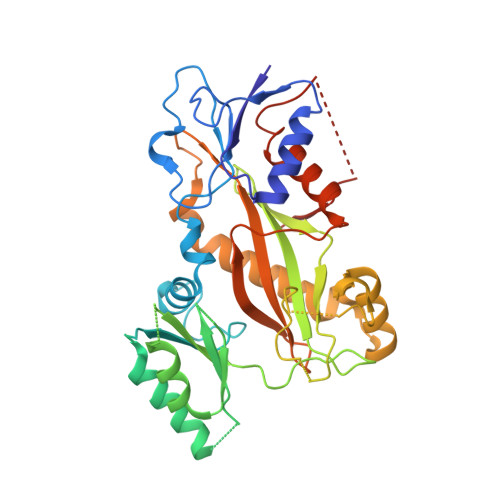
Structural insight into the stabilization of microtubules by taxanes.
Prota, A.E., Lucena-Agell, D., Ma, Y., Estevez-Gallego, J., Li, S., Bargsten, K., Josa-Prado, F., Altmann, K.H., Gaillard, N., Kamimura, S., Muhlethaler, T., Gago, F., Oliva, M.A., Steinmetz, M.O., Fang, W.S., Diaz, J.F.(2023) Elife 12
- PubMed: 36876916 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.7554/eLife.84791
- Primary Citation Related Structures:
8BDE, 8BDF, 8BDG - PubMed Abstract:
Paclitaxel (Taxol) is a taxane and a chemotherapeutic drug that stabilizes microtubules. While the interaction of paclitaxel with microtubules is well described, the lack of high-resolution structural information on a tubulin-taxane complex precludes a comprehensive description of the binding determinants that affect its mechanism of action. Here, we solved the crystal structure of baccatin III the core moiety of paclitaxel-tubulin complex at 1.9 Å resolution. Based on this information, we engineered taxanes with modified C13 side chains, solved their crystal structures in complex with tubulin, and analyzed their effects on microtubules (X-ray fiber diffraction), along with those of paclitaxel, docetaxel, and baccatin III. Further comparison of high-resolution structures and microtubules' diffractions with the apo forms and molecular dynamics approaches allowed us to understand the consequences of taxane binding to tubulin in solution and under assembled conditions. The results sheds light on three main mechanistic questions: (1) taxanes bind better to microtubules than to tubulin because tubulin assembly is linked to a βM-loopconformational reorganization (otherwise occludes the access to the taxane site) and, bulky C13 side chains preferentially recognize the assembled conformational state; (2) the occupancy of the taxane site has no influence on the straightness of tubulin protofilaments and; (3) longitudinal expansion of the microtubule lattices arises from the accommodation of the taxane core within the site, a process that is no related to the microtubule stabilization (baccatin III is biochemically inactive). In conclusion, our combined experimental and computational approach allowed us to describe the tubulin-taxane interaction in atomic detail and assess the structural determinants for binding.
- Laboratory of Biomolecular Research, Division of Biology and Chemistry, Paul Scherrer Institute, Villigen PSI, Switzerland.
Organizational Affiliation: