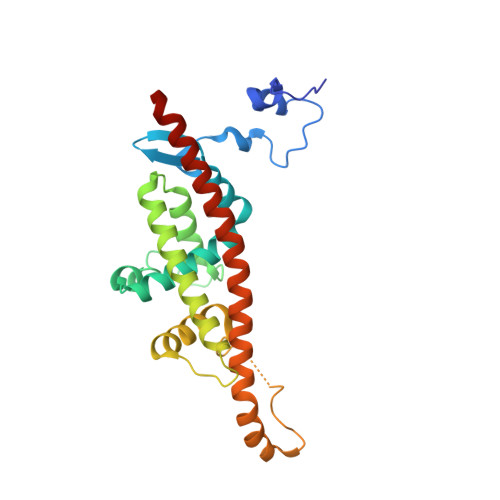
Atomic structure of a nudivirus occlusion body protein determined from a 70-year-old crystal sample.
Keown, J.R., Crawshaw, A.D., Trincao, J., Carrique, L., Gildea, R.J., Horrell, S., Warren, A.J., Axford, D., Owen, R., Evans, G., Bezier, A., Metcalf, P., Grimes, J.M.(2023) Nat Commun 14: 4160-4160
- PubMed: 37443157 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-023-39819-1
- Primary Citation Related Structures:
8BBT, 8BC5, 8BCK, 8BCL - PubMed Abstract:
Infectious protein crystals are an essential part of the viral lifecycle for double-stranded DNA Baculoviridae and double-stranded RNA cypoviruses. These viral protein crystals, termed occlusion bodies or polyhedra, are dense protein assemblies that form a crystalline array, encasing newly formed virions. Here, using X-ray crystallography we determine the structure of a polyhedrin from Nudiviridae. This double-stranded DNA virus family is a sister-group to the baculoviruses, whose members were thought to lack occlusion bodies. The 70-year-old sample contains a well-ordered lattice formed by a predominantly α-helical building block that assembles into a dense, highly interconnected protein crystal. The lattice is maintained by extensive hydrophobic and electrostatic interactions, disulfide bonds, and domain switching. The resulting lattice is resistant to most environmental stresses. Comparison of this structure to baculovirus or cypovirus polyhedra shows a distinct protein structure, crystal space group, and unit cell dimensions, however, all polyhedra utilise common principles of occlusion body assembly.
- Division of Structural Biology, Wellcome Centre for Human Genetics, University of Oxford, Oxford, UK. Jeremy@strubi.ox.ac.uk.
Organizational Affiliation: