Synthesis of Uronic Acid 1-Azasugars as Putative Inhibitors of alpha-Iduronidase, beta-Glucuronidase and Heparanase.
Doherty, G.G., Ler, G.J.M., Wimmer, N., Bernhardt, P.V., Ashmus, R.A., Vocadlo, D.J., Armstrong, Z., Davies, G.J., Maccarana, M., Li, J.P., Kayal, Y., Ferro, V.(2023) Chembiochem 24: e202200619-e202200619
- PubMed: 36453606 Search on PubMed
- DOI: https://doi.org/10.1002/cbic.202200619
- Primary Citation Related Structures:
8BAC - PubMed Abstract:
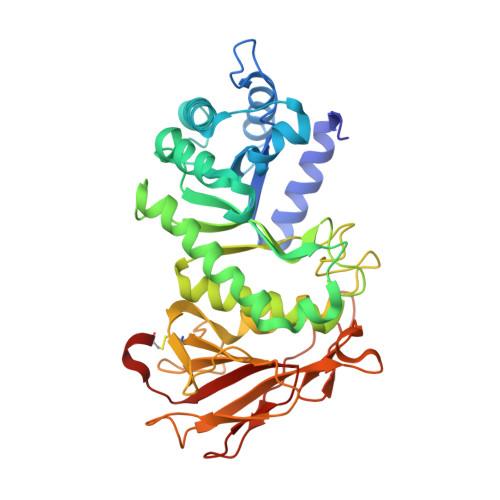
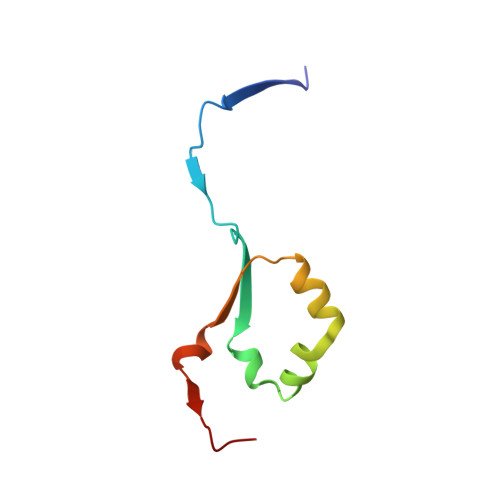
1-Azasugar analogues of l-iduronic acid (l-IdoA) and d-glucuronic acid (d-GlcA) and their corresponding enantiomers have been synthesized as potential pharmacological chaperones for mucopolysaccharidosis I (MPS I), a lysosomal storage disease caused by mutations in the gene encoding α-iduronidase (IDUA). The compounds were efficiently synthesized in nine or ten steps from d- or l-arabinose, and the structures were confirmed by X-ray crystallographic analysis of key intermediates. All compounds were inactive against IDUA, although l-IdoA-configured 8 moderately inhibited β-glucuronidase (β-GLU). The d-GlcA-configured 9 was a potent inhibitor of β-GLU and a moderate inhibitor of the endo-β-glucuronidase heparanase. Co-crystallization of 9 with heparanase revealed that the endocyclic nitrogen of 9 forms close interactions with both the catalytic acid and catalytic nucleophile.
- School of Chemistry & Molecular Biosciences, The University of Queensland, Brisbane, Queensland, 4072, Australia.
Organizational Affiliation: