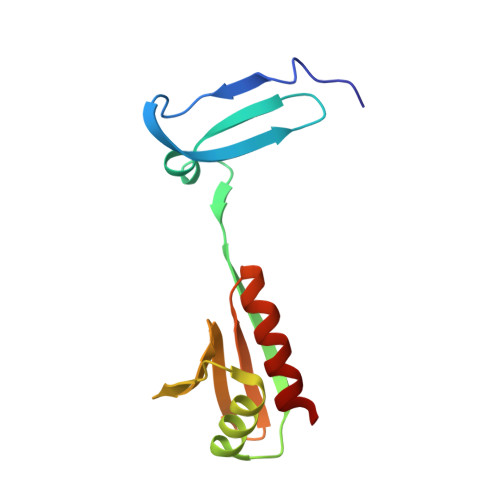
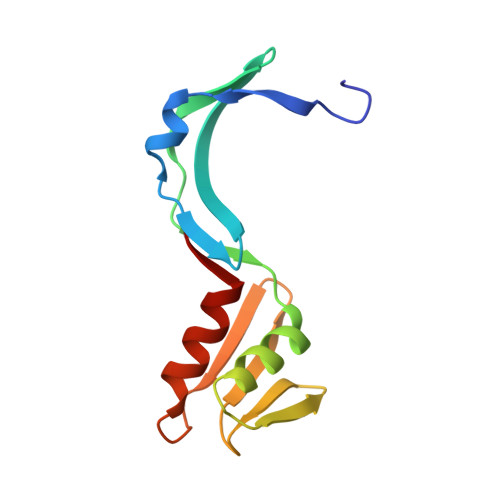
Molecular details of the CPSF73-CPSF100 C-terminal heterodimer and interaction with Symplekin.
Thore, S., Raoelijaona, F., Talenton, V., Fribourg, S., Mackereth, C.D.(2023) Open Biol 13: 230221-230221
- PubMed: 37989222 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1098/rsob.230221
- Primary Citation Related Structures:
8B7T, 8BA1 - PubMed Abstract:
Eukaryotic pre-mRNA is processed by a large multiprotein complex to accurately cleave the 3' end, and to catalyse the addition of the poly(A) tail. Within this cleavage and polyadenylation specificity factor (CPSF) machinery, the CPSF73/CPSF3 endonuclease subunit directly contacts both CPSF100/CPSF2 and the scaffold protein Symplekin to form a subcomplex known as the core cleavage complex or mammalian cleavage factor. Here we have taken advantage of a stable CPSF73-CPSF100 minimal heterodimer from Encephalitozoon cuniculi to determine the solution structure formed by the first and second C-terminal domain (CTD1 and CTD2) of both proteins. We find a large number of contacts between both proteins in the complex, and notably in the region between CTD1 and CTD2. A similarity is also observed between CTD2 and the TATA-box binding protein (TBP) domains. Separately, we have determined the structure of the terminal CTD3 domain of CPSF73, which also belongs to the TBP domain family and is connected by a flexible linker to the rest of CPSF73. Biochemical assays demonstrate a key role for the CTD3 of CPSF73 in binding Symplekin, and structural models of the trimeric complex from other species allow for comparative analysis and support an overall conserved architecture.
- Inserm, CNRS, ARNA Laboratory, Univ. Bordeaux, U1212, UMR 5320, 33000 Bordeaux, France.
Organizational Affiliation: