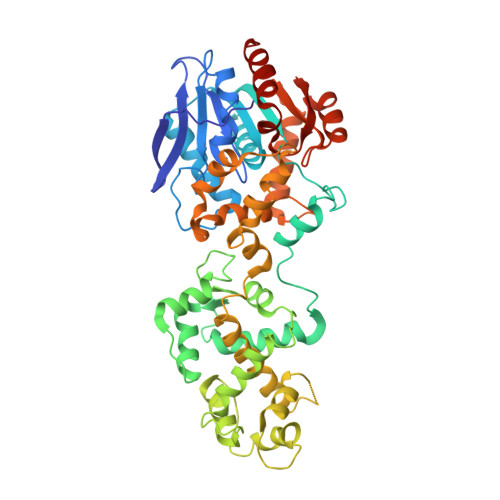
X-ray structure of the haloalkane dehalogenase HaloTag7 with an insertion of Calmodulin-M13 fusion at position 154-156 that mimic the structure of CaProLa, an calcium gated protein labeling technology
Tarnawski, M., Johnsson, K., Hiblot, J.To be published.