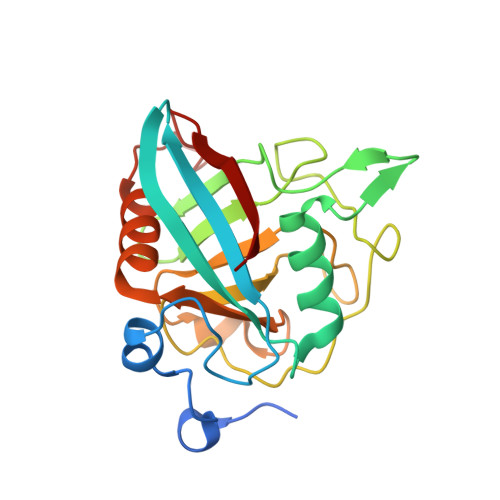
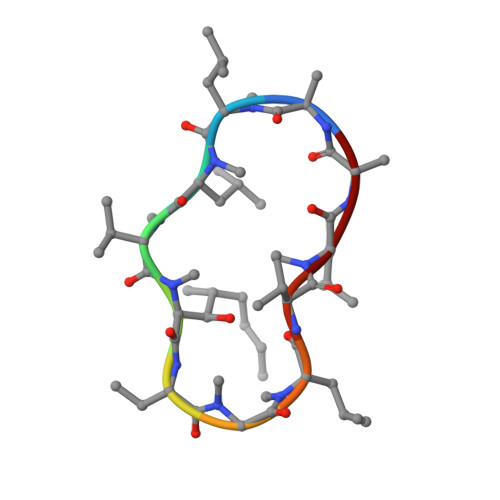
Structural Basis for Cyclosporin Isoform-Specific Inhibition of Cyclophilins from Toxoplasma gondii .
Favretto, F., Jimenez-Faraco, E., Conter, C., Dominici, P., Hermoso, J.A., Astegno, A.(2023) ACS Infect Dis 9: 365-377
- PubMed: 36653744 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/acsinfecdis.2c00566
- Primary Citation Related Structures:
8B58 - PubMed Abstract:
Cyclosporin (CsA) has antiparasite activity against the human pathogen Toxoplasma gondii . A possible mechanism of action involves CsA binding to T. gondii cyclophilins, although much remains to be understood. Herein, we characterize the functional and structural properties of a conserved (TgCyp23) and a more divergent (TgCyp18.4) cyclophilin isoform from T. gondii . While TgCyp23 is a highly active cis-trans-prolyl isomerase (PPIase) and binds CsA with nanomolar affinity, TgCyp18.4 shows low PPIase activity and is significantly less sensitive to CsA inhibition. The crystal structure of the TgCyp23:CsA complex was solved at the atomic resolution showing the molecular details of CsA recognition by the protein. Computational and structural studies revealed relevant differences at the CsA-binding site between TgCyp18.4 and TgCyp23, suggesting that the two cyclophilins might have distinct functions in the parasite. These studies highlight the extensive diversification of TgCyps and pave the way for antiparasite interventions based on selective targeting of cyclophilins.
- Department of Biotechnology, University of Verona, Strada Le Grazie 15, 37134Verona, Italy.
Organizational Affiliation: