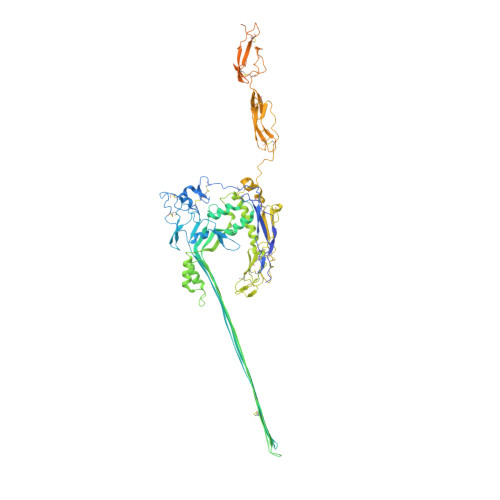
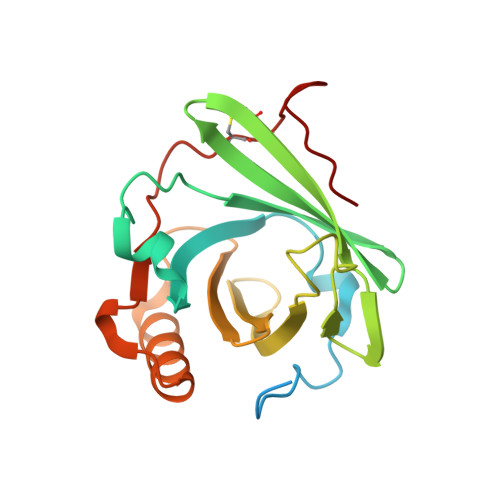
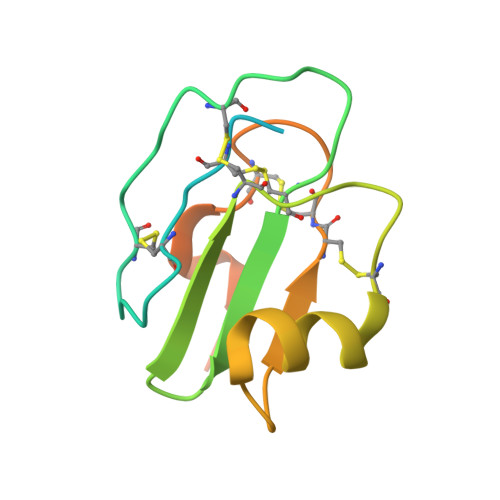
Structural basis for membrane attack complex inhibition by CD59.
Couves, E.C., Gardner, S., Voisin, T.B., Bickel, J.K., Stansfeld, P.J., Tate, E.W., Bubeck, D.(2023) Nat Commun 14: 890-890
- PubMed: 36797260 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-023-36441-z
- Primary Citation Related Structures:
8B0F, 8B0G, 8B0H - PubMed Abstract:
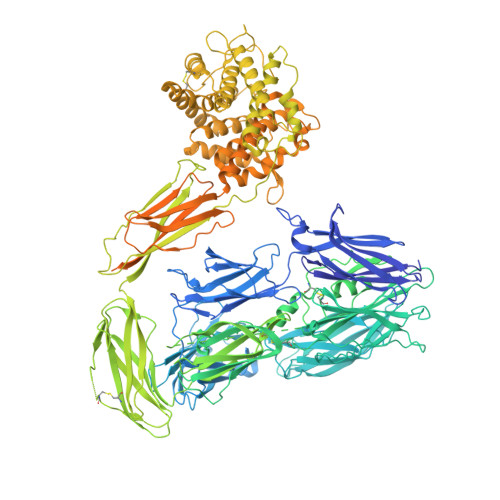
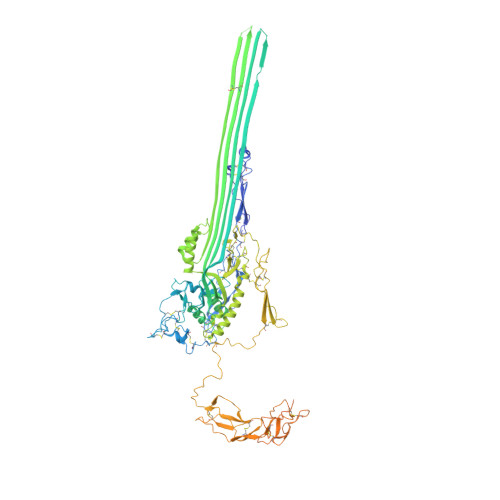
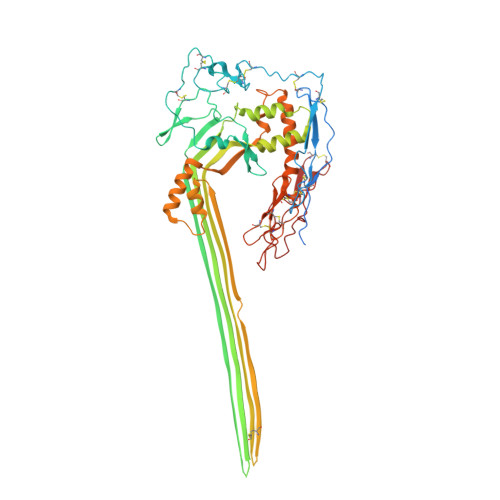
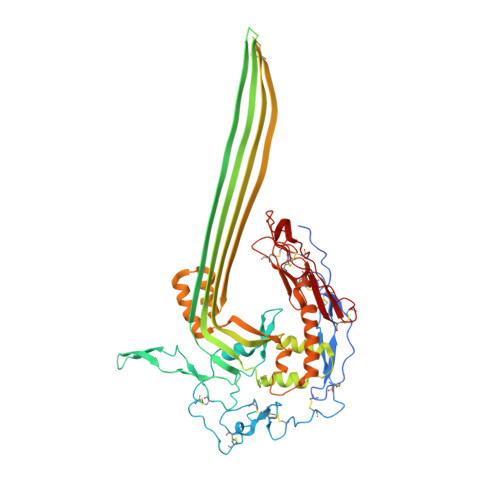
CD59 is an abundant immuno-regulatory receptor that protects human cells from damage during complement activation. Here we show how the receptor binds complement proteins C8 and C9 at the membrane to prevent insertion and polymerization of membrane attack complex (MAC) pores. We present cryo-electron microscopy structures of two inhibited MAC precursors known as C5b8 and C5b9. We discover that in both complexes, CD59 binds the pore-forming β-hairpins of C8 to form an intermolecular β-sheet that prevents membrane perforation. While bound to C8, CD59 deflects the cascading C9 β-hairpins, rerouting their trajectory into the membrane. Preventing insertion of C9 restricts structural transitions of subsequent monomers and indirectly halts MAC polymerization. We combine our structural data with cellular assays and molecular dynamics simulations to explain how the membrane environment impacts the dual roles of CD59 in controlling pore formation of MAC, and as a target of bacterial virulence factors which hijack CD59 to lyse human cells.
- Department of Life Sciences, Sir Ernst Chain Building, Imperial College London, London, SW7 2AZ, United Kingdom.
Organizational Affiliation: