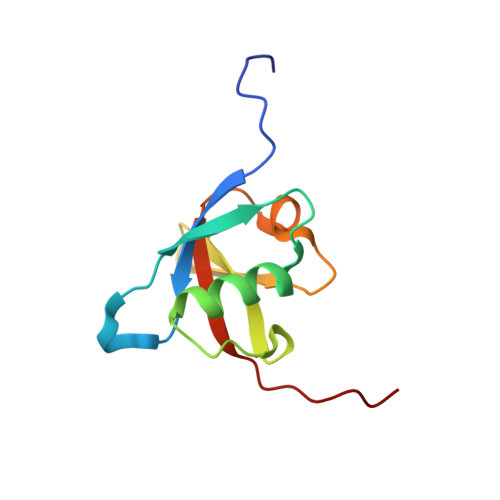
Structural insights into the complex of oncogenic KRas4B G12V and Rgl2, a RalA/B activator.
Tariq, M., Ikeya, T., Togashi, N., Fairall, L., Kamei, S., Mayooramurugan, S., Abbott, L.R., Hasan, A., Bueno-Alejo, C., Sukegawa, S., Romartinez-Alonso, B., Muro Campillo, M.A., Hudson, A.J., Ito, Y., Schwabe, J.W., Dominguez, C., Tanaka, K.(2024) Life Sci Alliance 7
- PubMed: 37833074 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.26508/lsa.202302080
- Primary Citation Related Structures:
8AU4, 8B69 - PubMed Abstract:
About a quarter of total human cancers carry mutations in Ras isoforms. Accumulating evidence suggests that small GTPases, RalA, and RalB, and their activators, Ral guanine nucleotide exchange factors (RalGEFs), play an essential role in oncogenic Ras-induced signalling. We studied the interaction between human KRas4B and the Ras association (RA) domain of Rgl2 (Rgl2 RA ), one of the RA-containing RalGEFs. We show that the G12V oncogenic KRas4B mutation changes the interaction kinetics with Rgl2 RA The crystal structure of the KRas4B G12V : Rgl2 RA complex shows a 2:2 heterotetramer where the switch I and switch II regions of each KRas G12V interact with both Rgl2 RA molecules. This structural arrangement is highly similar to the HRas E31K :RALGDS RA crystal structure and is distinct from the well-characterised Ras:Raf complex. Interestingly, the G12V mutation was found at the dimer interface of KRas4B G12V with its partner. Our study reveals a potentially distinct mode of Ras:effector complex formation by RalGEFs and offers a possible mechanistic explanation for how the oncogenic KRas4B G12V hyperactivates the RalA/B pathway.
- https://ror.org/04h699437 Department of Molecular and Cell Biology, University of Leicester, Leicester, UK.
Organizational Affiliation: