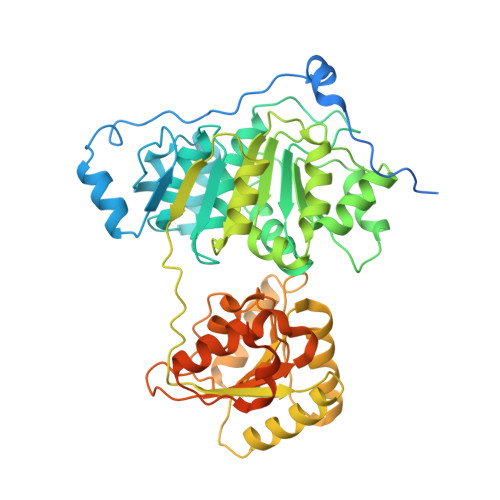
Nonstructural N- and C-tails of Dbp2 confer the protein full helicase activities.
Song, Q.X., Liu, N.N., Liu, Z.X., Zhang, Y.Z., Rety, S., Hou, X.M., Xi, X.G.(2023) J Biological Chem 299: 104592-104592
- PubMed: 36894019 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jbc.2023.104592
- Primary Citation Related Structures:
8ARK, 8ARP - PubMed Abstract:
Human DDX5 and its yeast ortholog Dbp2 are ATP-dependent RNA helicases that play a key role in normal cell processes, cancer development, and viral infection. The crystal structure of the RecA1-like domain of DDX5 is available but the global structure of DDX5/Dbp2 subfamily proteins remains to be elucidated. Here, we report the first X-ray crystal structures of the Dbp2 helicase core alone and in complex with ADP at 3.22 Å and 3.05 Å resolutions, respectively. The structures of the ADP-bound post-hydrolysis state and apo-state demonstrate the conformational changes that occur when the nucleotides are released. Our results showed that the helicase core of Dbp2 shifted between open and closed conformation in solution but the unwinding activity was hindered when the helicase core was restricted to a single conformation. A small-angle X-ray scattering experiment showed that the disordered amino (N) tail and carboxy (C) tails are flexible in solution. Truncation mutations confirmed that the terminal tails were critical for the nucleic acid binding, ATPase, and unwinding activities, with the C-tail being exclusively responsible for the annealing activity. Furthermore, we labeled the terminal tails to observe the conformational changes between the disordered tails and the helicase core upon binding nucleic acid substrates. Specifically, we found that the nonstructural terminal tails bind to RNA substrates and tether them to the helicase core domain, thereby conferring full helicase activities to the Dbp2 protein. This distinct structural characteristic provides new insight into the mechanism of DEAD-box RNA helicases.
- College of Life Sciences, State Key Laboratory of Crop Stress Biology in Arid Areas, Northwest A&F University, Yangling, Shaanxi, PR China.
Organizational Affiliation: