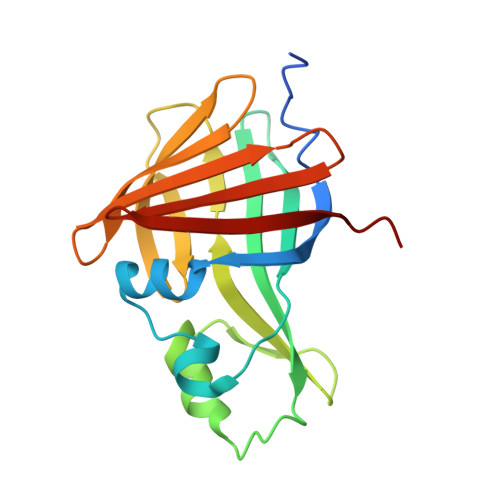
Illuminating the mechanism and allosteric behavior of NanoLuc luciferase.
Nemergut, M., Pluskal, D., Horackova, J., Sustrova, T., Tulis, J., Barta, T., Baatallah, R., Gagnot, G., Novakova, V., Majerova, M., Sedlackova, K., Marques, S.M., Toul, M., Damborsky, J., Prokop, Z., Bednar, D., Janin, Y.L., Marek, M.(2023) Nat Commun 14: 7864-7864
- PubMed: 38030625 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-023-43403-y
- Primary Citation Related Structures:
8AQ6, 8AQH, 8AQI, 8BO9 - PubMed Abstract:
NanoLuc, a superior β-barrel fold luciferase, was engineered 10 years ago but the nature of its catalysis remains puzzling. Here experimental and computational techniques are combined, revealing that imidazopyrazinone luciferins bind to an intra-barrel catalytic site but also to an allosteric site shaped on the enzyme surface. Structurally, binding to the allosteric site prevents simultaneous binding to the catalytic site, and vice versa, through concerted conformational changes. We demonstrate that restructuration of the allosteric site can boost the luminescent reaction in the remote active site. Mechanistically, an intra-barrel arginine coordinates the imidazopyrazinone component of luciferin, which reacts with O 2 via a radical charge-transfer mechanism, and then it also protonates the resulting excited amide product to form a light-emitting neutral species. Concomitantly, an aspartate, supported by two tyrosines, fine-tunes the blue color emitter to secure a high emission intensity. This information is critical to engineering the next-generation of ultrasensitive bioluminescent reporters.
- Loschmidt Laboratories, Department of Experimental Biology and RECETOX, Faculty of Science, Masaryk University, Kamenice 5, Bld. C13, 625 00, Brno, Czech Republic.
Organizational Affiliation: