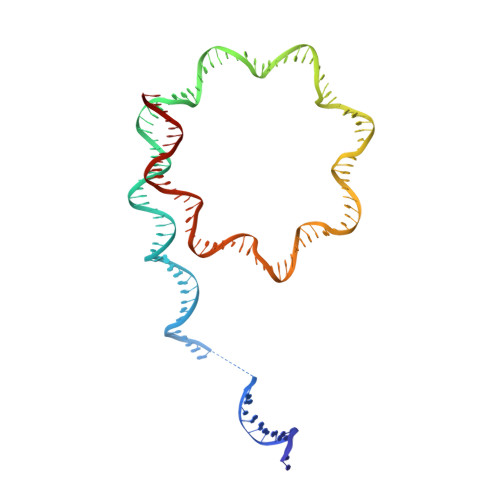
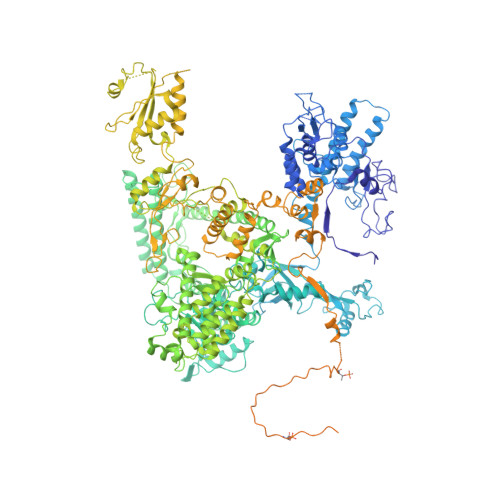
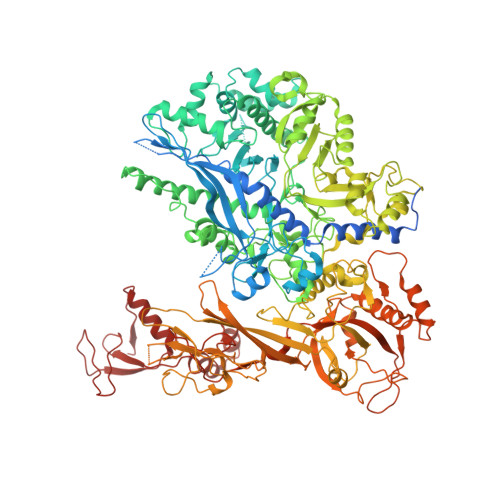
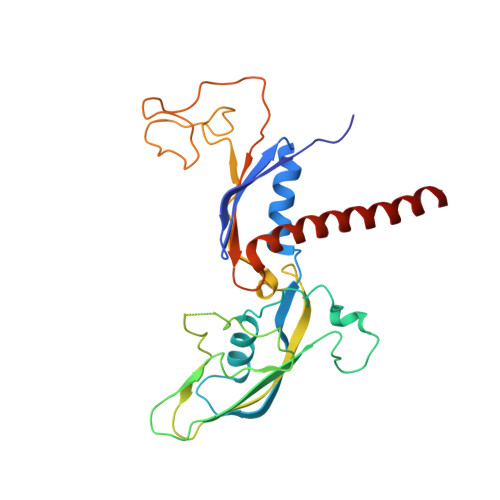
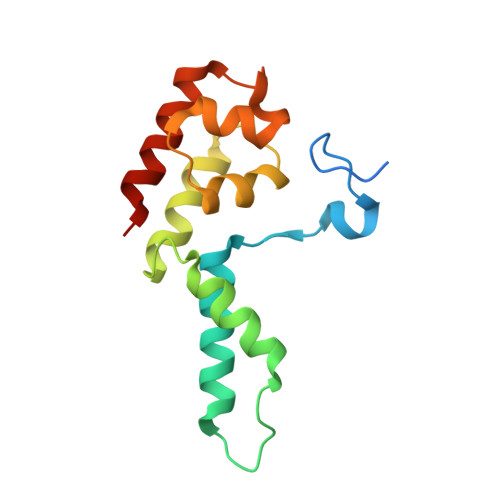
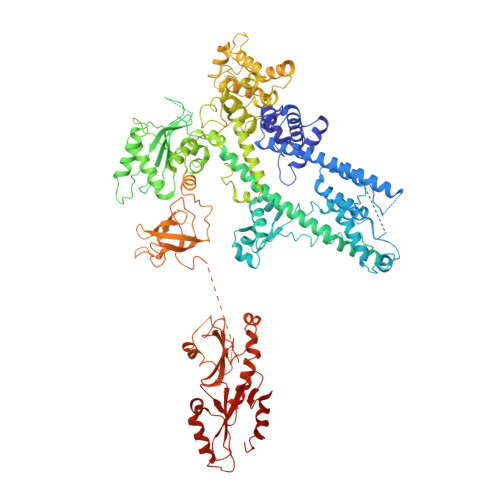
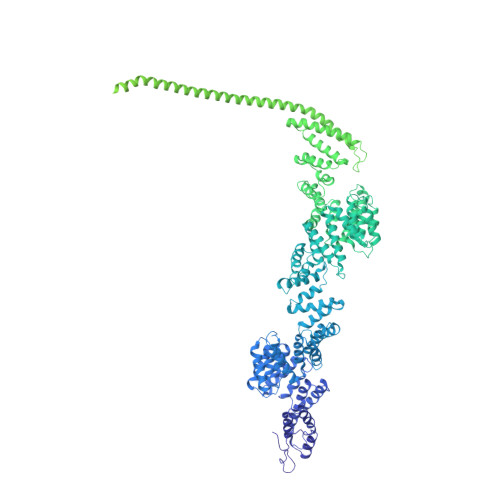
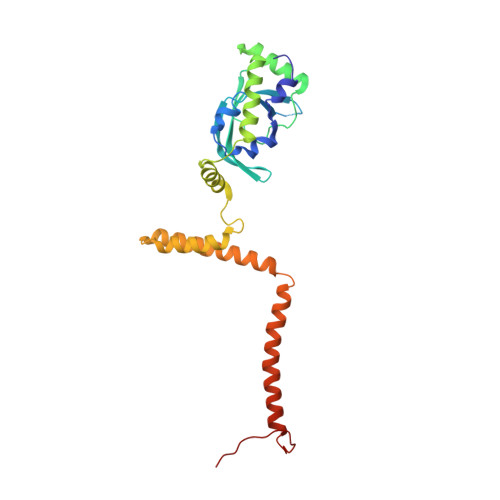
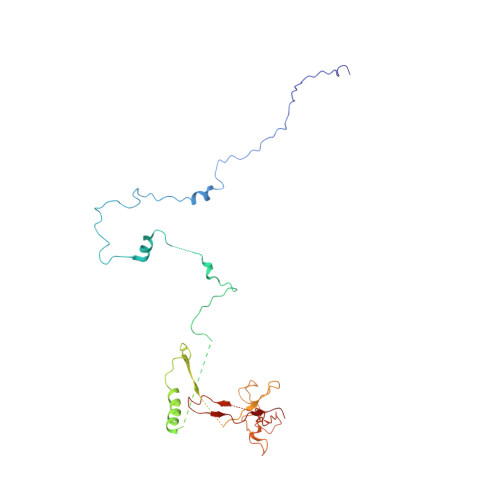
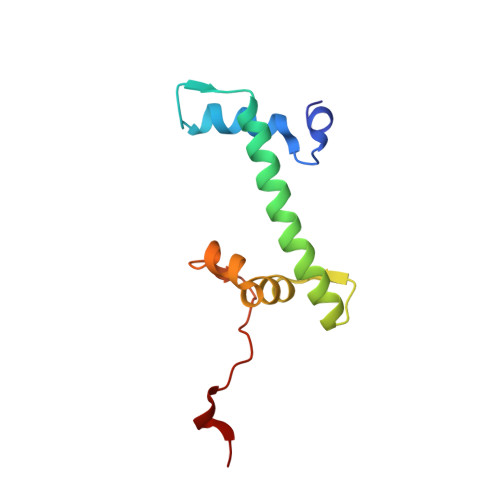
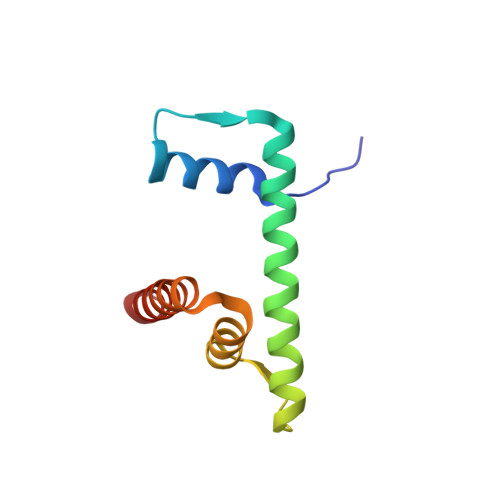
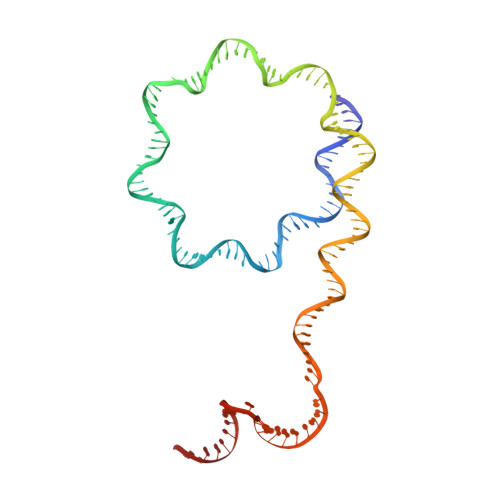
Structure of a backtracked hexasomal intermediate of nucleosome transcription.
Farnung, L., Ochmann, M., Garg, G., Vos, S.M., Cramer, P.(2022) Mol Cell 82: 3126-3134.e7
- PubMed: 35858621 Search on PubMed
- DOI: https://doi.org/10.1016/j.molcel.2022.06.027
- Primary Citation Related Structures:
8A3Y, 8A40 - PubMed Abstract:
During gene transcription, RNA polymerase II (RNA Pol II) passes nucleosomes with the help of various elongation factors. Here, we show that RNA Pol II achieves efficient nucleosome passage when the human elongation factors DSIF, PAF1 complex (PAF), RTF1, SPT6, and TFIIS are present. The cryo-EM structure of an intermediate of the nucleosome passage shows a partially unraveled hexasome that lacks the proximal H2A-H2B dimer and interacts with the RNA Pol II jaw, DSIF, and the CTR9trestle helix. RNA Pol II adopts a backtracked state with the RNA 3' end dislodged from the active site and bound in the RNA Pol II pore. Additional structures and biochemical data show that human TFIIS enters the RNA Pol II pore and stimulates the cleavage of the backtracked RNA and nucleosome passage.
- Max Planck Institute for Multidisciplinary Sciences, Department of Molecular Biology, Am Fassberg 11, 37077 Göttingen, Germany. Electronic address: lucas_farnung@hms.harvard.edu.
Organizational Affiliation: