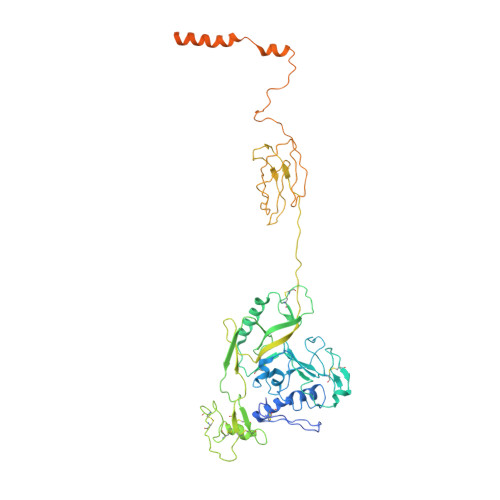
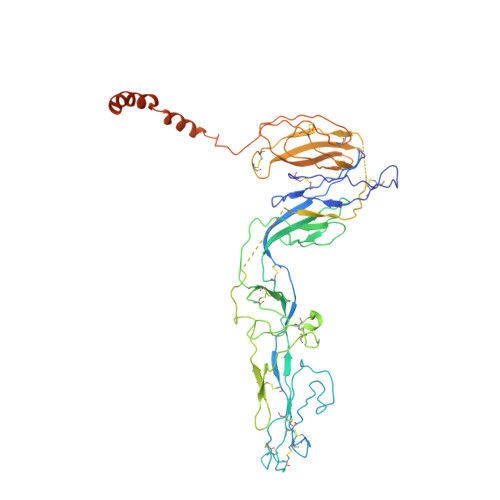
Architecture of severe fever with thrombocytopenia syndrome virus.
Sun, Z., Cheng, J., Bai, Y., Cao, L., Xie, D., Deng, F., Zhang, X., Rao, Z., Lou, Z.(2023) Protein Cell 14: 914-918
- PubMed: 37038326 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/procel/pwad019
- Primary Citation Related Structures:
7X6U, 7X6W, 7X72 - Department of Basic Research, Guangzhou Laboratory, Guangzhou 510005, China.
Organizational Affiliation: