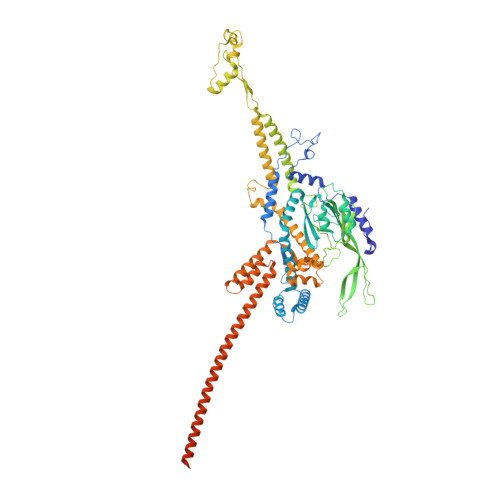
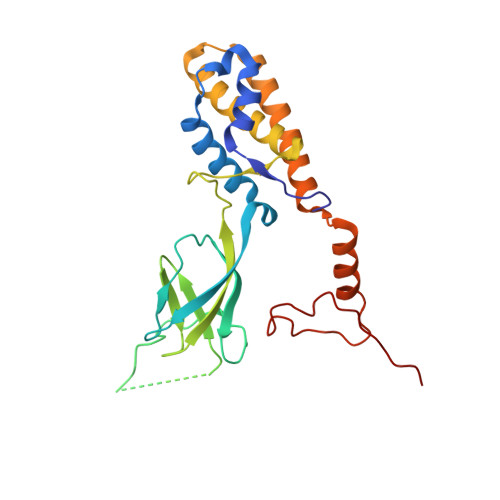
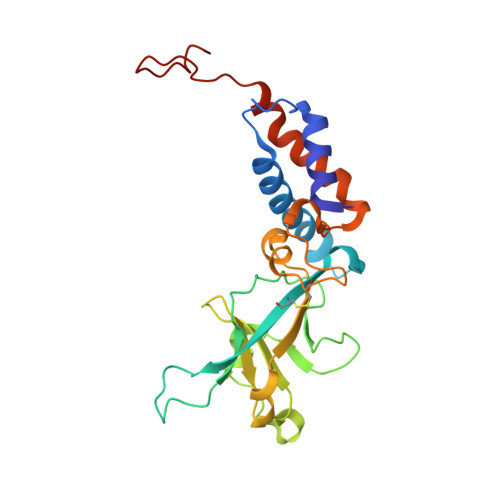
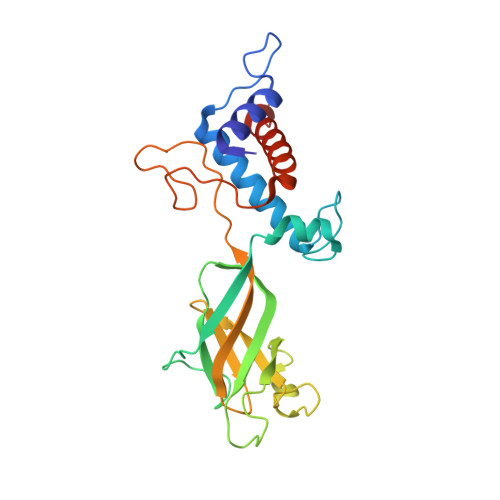
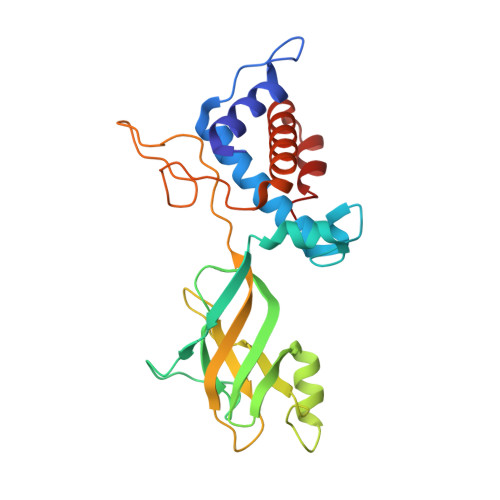
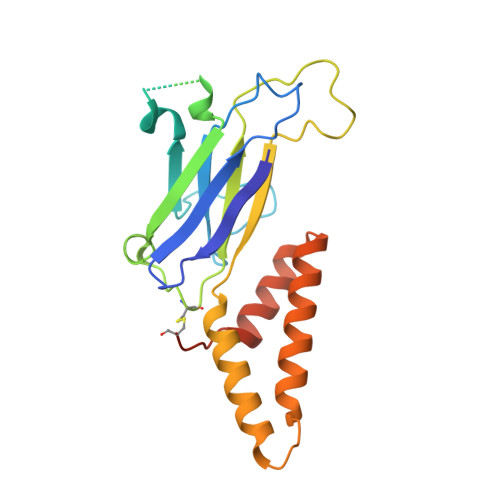
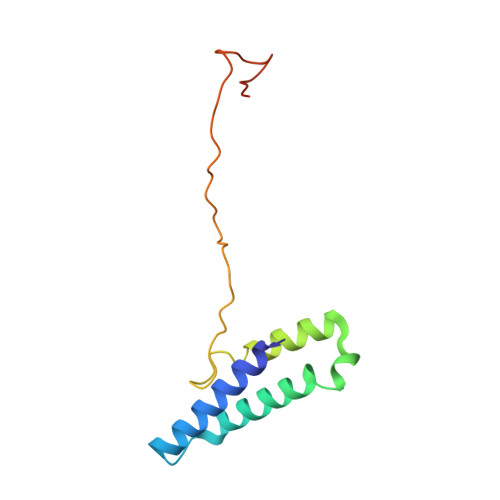
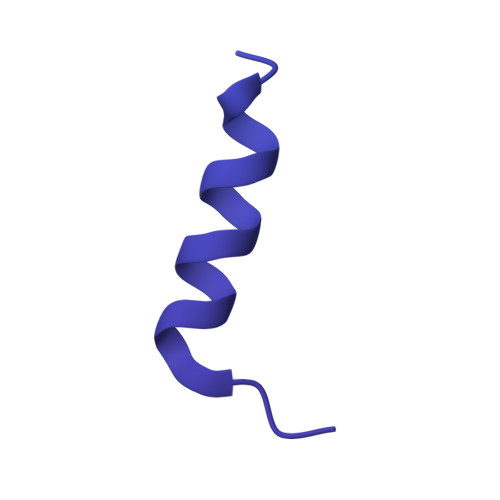
Structural atlas of a human gut crassvirus.
Bayfield, O.W., Shkoporov, A.N., Yutin, N., Khokhlova, E.V., Smith, J.L.R., Hawkins, D.E.D.P., Koonin, E.V., Hill, C., Antson, A.A.(2023) Nature 617: 409-416
- PubMed: 37138077 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41586-023-06019-2
- Primary Citation Related Structures:
7QOF, 7QOG, 7QOH, 7QOI, 7QOJ, 7QOK, 7QOL - PubMed Abstract:
CrAssphage and related viruses of the order Crassvirales (hereafter referred to as crassviruses) were originally discovered by cross-assembly of metagenomic sequences. They are the most abundant viruses in the human gut, are found in the majority of individual gut viromes, and account for up to 95% of the viral sequences in some individuals 1-4 . Crassviruses are likely to have major roles in shaping the composition and functionality of the human microbiome, but the structures and roles of most of the virally encoded proteins are unknown, with only generic predictions resulting from bioinformatic analyses 4,5 . Here we present a cryo-electron microscopy reconstruction of Bacteroides intestinalis virus ΦcrAss001 6 , providing the structural basis for the functional assignment of most of its virion proteins. The muzzle protein forms an assembly about 1 MDa in size at the end of the tail and exhibits a previously unknown fold that we designate the 'crass fold', that is likely to serve as a gatekeeper that controls the ejection of cargos. In addition to packing the approximately 103 kb of virus DNA, the ΦcrAss001 virion has extensive storage space for virally encoded cargo proteins in the capsid and, unusually, within the tail. One of the cargo proteins is present in both the capsid and the tail, suggesting a general mechanism for protein ejection, which involves partial unfolding of proteins during their extrusion through the tail. These findings provide a structural basis for understanding the mechanisms of assembly and infection of these highly abundant crassviruses.
- York Structural Biology Laboratory, Department of Chemistry, University of York, York, UK. oliver.bayfield@york.ac.uk.
Organizational Affiliation: