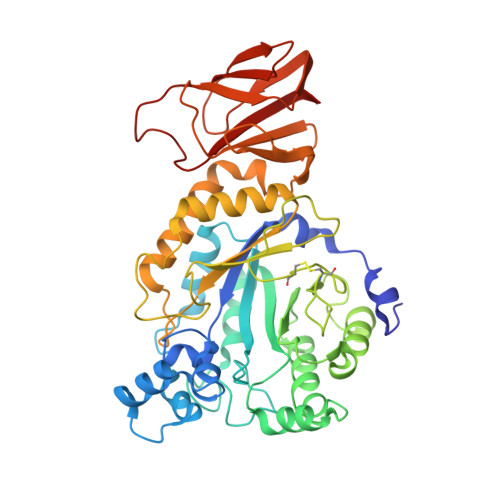
Cryo-EM structures of human fucosidase FucA1 reveal insight into substrate recognition and catalysis.
Armstrong, Z., Meek, R.W., Wu, L., Blaza, J.N., Davies, G.J.(2022) Structure 30: 1443
- PubMed: 35907402 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.str.2022.07.001
- Primary Citation Related Structures:
7PLS, 7PM4 - PubMed Abstract:
Enzymatic hydrolysis of α-L-fucose from fucosylated glycoconjugates is consequential in bacterial infections and the neurodegenerative lysosomal storage disorder fucosidosis. Understanding human α-L-fucosidase catalysis, in an effort toward drug design, has been hindered by the absence of three-dimensional structural data for any animal fucosidase. Here, we have used cryoelectron microscopy (cryo-EM) to determine the structure of human lysosomal α-L-fucosidase (FucA1) in both an unliganded state and in complex with the inhibitor deoxyfuconojirimycin. These structures, determined at 2.49 Å resolution, reveal the homotetrameric structure of FucA1, the architecture of the catalytic center, and the location of both natural population variations and disease-causing mutations. Furthermore, this work has conclusively identified the hitherto contentious identity of the catalytic acid/base as aspartate-276, representing a shift from both the canonical glutamate acid/base residue and a previously proposed glutamate residue. These findings have furthered our understanding of how FucA1 functions in both health and disease.
- Department of Chemistry, Structural Biology Laboratory, University of York, Heslington, York YO10 5DD, UK.
Organizational Affiliation: