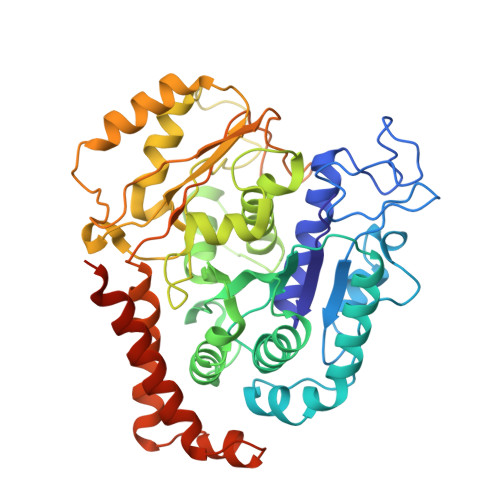
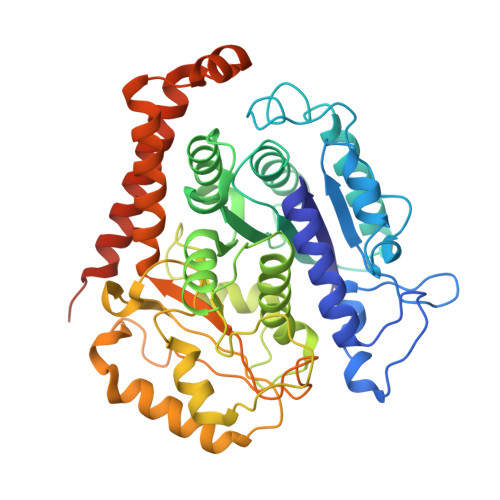
Conformational changes in tubulin upon binding cryptophycin-52 reveal its mechanism of action.
Eren, E., Watts, N.R., Sackett, D.L., Wingfield, P.T.(2021) J Biological Chem 297: 101138-101138
- PubMed: 34461087 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jbc.2021.101138
- Primary Citation Related Structures:
7LXB, 7M18, 7M20 - PubMed Abstract:
Cryptophycin-52 (Cp-52) is potentially the most potent anticancer drug known, with IC 50 values in the low picomolar range, but its binding site on tubulin and mechanism of action are unknown. Here, we have determined the binding site of Cp-52, and its parent compound, cryptophycin-1, on HeLa tubulin, to a resolution of 3.3 Å and 3.4 Å, respectively, by cryo-EM and characterized this binding further by molecular dynamics simulations. The binding site was determined to be located at the tubulin interdimer interface and partially overlap that of maytansine, another cytotoxic tubulin inhibitor. Binding induces curvature both within and between tubulin dimers that is incompatible with the microtubule lattice. Conformational changes occur in both α-tubulin and β-tubulin, particularly in helices H8 and H10, with distinct differences between α and β monomers and between Cp-52-bound and cryptophycin-1-bound tubulin. From these results, we have determined: (i) the mechanism of action of inhibition of both microtubule polymerization and depolymerization, (ii) how the affinity of Cp-52 for tubulin may be enhanced, and (iii) where linkers for targeted delivery can be optimally attached to this molecule.
- Protein Expression Laboratory, NIAMS, National Institutes of Health, Bethesda, Maryland, USA.
Organizational Affiliation: