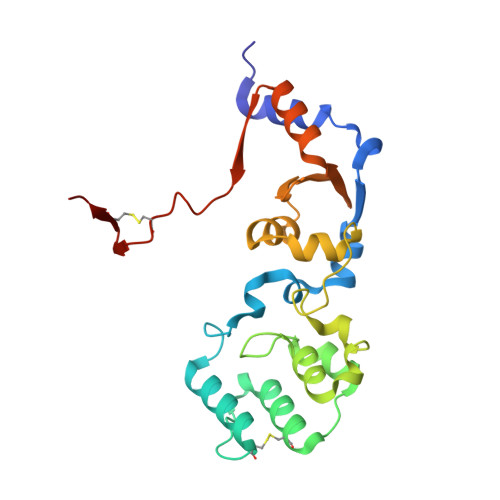
Sensor Domain of Histidine Kinase VxrA of Vibrio cholerae - A Hairpin-swapped Dimer and its Conformational Change.
Tan, K., Teschler, J.K., Wu, R., Jedrzejczak, R.P., Zhou, M., Shuvalova, L.A., Endres, M.J., Welk, L.F., Kwon, K., Anderson, W.F., Satchell, K.J.F., Yildiz, F.H., Joachimiak, A.(2021) J Bacteriol
- PubMed: 33753465 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1128/JB.00643-20
- Primary Citation Related Structures:
4R7Q, 7KB3, 7KB7, 7KB9, 7LA6 - PubMed Abstract:
VxrA and VxrB are cognate histidine kinase (HK) - response regulator (RR) pairs of a two-component signaling system (TCS) found in Vibrio cholerae , a bacterial pathogen that causes cholera. The VxrAB TCS positively regulates virulence, the Type VI Secretion System, biofilm formation, and cell wall homeostasis in V. cholerae , providing protection from environmental stresses and contributing to the transmission and virulence of the pathogen. The VxrA HK has a unique periplasmic sensor domain (SD) and, remarkably, lacks a cytoplasmic linker domain between the second transmembrane helix and the dimerization and histidine phosphotransfer (DHp) domain, indicating that this system may utilize a potentially unique signal sensing and transmission TCS mechanism. In this study, we have determined several crystal structures of VxrA-SD and its mutants. These structures reveal a novel structural fold forming an unusual β hairpin-swapped dimer. A conformational change caused by relative rotation of the two monomers in a VxrA-SD dimer could potentially change the association of transmembrane helices and, subsequently, the pairing of cytoplasmic DHp domains. Based on the structural observation, we propose a putative scissor-like closing regulation mechanism for the VxrA HK. IMPORTANCE V. cholerae has a dynamic life cycle, which requires rapid adaptation to changing external conditions. Two-component signal transduction (TCS) systems allow V. cholerae to sense and respond to these environmental changes. The VxrAB TCS positively regulates a number of important V. cholerae phenotypes, including virulence, the Type Six Secretion System, biofilm formation, and cell wall homeostasis. Here, we provide the crystal structure of the VxrA sensor histidine kinase sensing domain and propose a mechanism for signal transduction. The cognate signal for VxrAB remains unknown, however, in this work we couple our structural analysis with functional assessments of key residues to further our understanding of this important TCS.
- Center for Structural Genomics of Infectious Diseases, University of Chicago, 5735 South Ellis Avenue, Chicago, IL 60637, USA.
Organizational Affiliation: