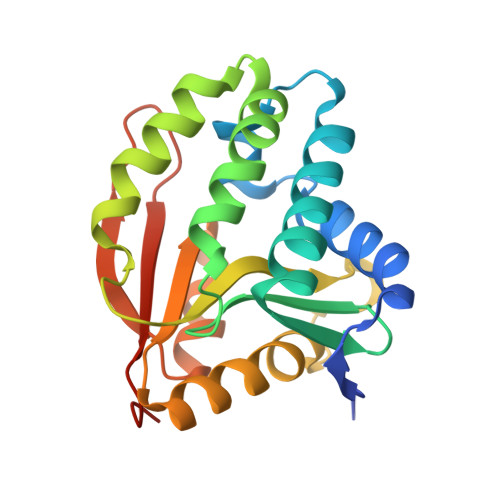
Structure-guided selection of puromycin N-acetyltransferase mutants with enhanced selection stringency for deriving mammalian cell lines expressing recombinant proteins.
Caputo, A.T., Eder, O.M., Bereznakova, H., Pothuis, H., Ardevol, A., Newman, J., Nuttall, S., Peat, T.S., Adams, T.E.(2021) Sci Rep 11: 5247-5247
- PubMed: 33664348 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41598-021-84551-9
- Primary Citation Related Structures:
7K09, 7K0A - PubMed Abstract:
Puromycin and the Streptomyces alboniger-derived puromycin N-acetyltransferase (PAC) enzyme form a commonly used system for selecting stably transfected cultured cells. The crystal structure of PAC has been solved using X-ray crystallography, revealing it to be a member of the GCN5-related N-acetyltransferase (GNAT) family of acetyltransferases. Based on structures in complex with acetyl-CoA or the reaction products CoA and acetylated puromycin, four classes of mutations in and around the catalytic site were designed and tested for activity. Single-residue mutations were identified that displayed a range of enzymatic activities, from complete ablation to enhanced activity relative to wild-type (WT) PAC. Cell pools of stably transfected HEK293 cells derived using two PAC mutants with attenuated activity, Y30F and A142D, were found to secrete up to three-fold higher levels of a soluble, recombinant target protein than corresponding pools derived with the WT enzyme. A third mutant, Y171F, appeared to stabilise the intracellular turnover of PAC, resulting in an apparent loss of selection stringency. Our results indicate that the structure-guided manipulation of PAC function can be utilised to enhance selection stringency for the derivation of mammalian cell lines secreting elevated levels of recombinant proteins.
- Biomedical Manufacturing, Commonwealth Scientific and Industrial Research Organisation, 343 Royal Parade, Parkville, Victoria, 3052, Australia. alex.caputo@csiro.au.
Organizational Affiliation: