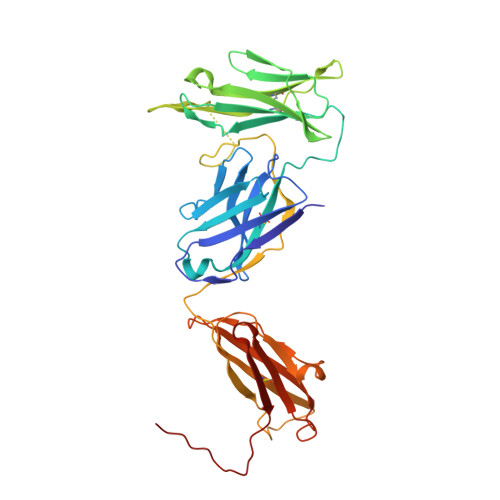
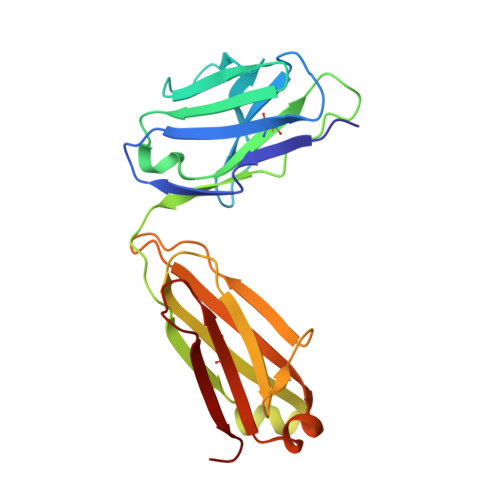
Structural basis of LAIR1 targeting by polymorphic Plasmodium RIFINs.
Xu, K., Wang, Y., Shen, C.H., Chen, Y., Zhang, B., Liu, K., Tsybovsky, Y., Wang, S., Farney, S.K., Gorman, J., Stephens, T., Verardi, R., Yang, Y., Zhou, T., Chuang, G.Y., Lanzavecchia, A., Piccoli, L., Kwong, P.D.(2021) Nat Commun 12: 4226-4226
- PubMed: 34244481 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-021-24291-6
- Primary Citation Related Structures:
7JZ1, 7JZ4, 7JZI, 7JZK - PubMed Abstract:
RIFIN, a large family of Plasmodium variant surface antigens, plays a crucial role in malaria pathogenesis by mediating immune suppression through activation of inhibitory receptors such as LAIR1, and antibodies with LAIR1 inserts have been identified that bind infected erythrocytes through RIFIN. However, details of RIFIN-mediated LAIR1 recognition and receptor activation have been unclear. Here, we use negative-stain EM to define the architecture of LAIR1-inserted antibodies and determine crystal structures of RIFIN-variable 2 (V2) domain in complex with a LAIR1 domain. These structures reveal the LAIR1-binding region of RIFIN to be hydrophobic and membrane-distal, to exhibit extensive structural diversity, and to interact with RIFIN-V2 in a one-to-one fashion. Through structural and sequence analysis of various LAIR1 constructs, we identify essential elements of RIFIN-binding on LAIR1. Furthermore, a structure-derived LAIR1-binding sequence signature ascertained >20 LAIR1-binding RIFINs, including some from P. falciparum field strains and Plasmodium species infecting gorillas and chimpanzees.
- Vaccine Research Center, National Institute of Allergy and Infectious Diseases, National Institutes of Health, Bethesda, MD, USA. xu.4692@osu.edu.
Organizational Affiliation: