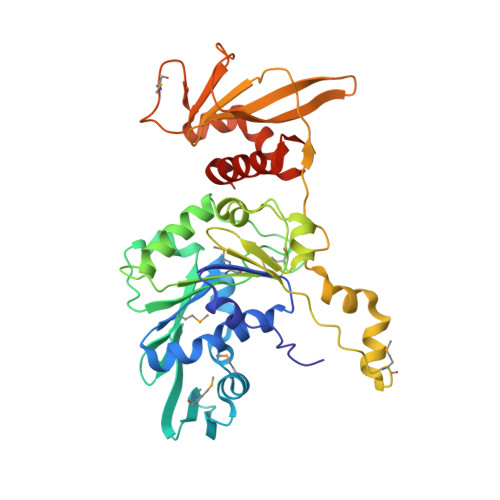
Atomistic basis of force generation, translocation, and coordination in a viral genome packaging motor.
Pajak, J., Dill, E., Reyes-Aldrete, E., White, M.A., Kelch, B.A., Jardine, P.J., Arya, G., Morais, M.C.(2021) Nucleic Acids Res 49: 6474-6488
- PubMed: 34050764 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gkab372
- Primary Citation Related Structures:
7JQ6, 7JQ7, 7JQP - PubMed Abstract:
Double-stranded DNA viruses package their genomes into pre-assembled capsids using virally-encoded ASCE ATPase ring motors. We present the first atomic-resolution crystal structure of a multimeric ring form of a viral dsDNA packaging motor, the ATPase of the asccφ28 phage, and characterize its atomic-level dynamics via long timescale molecular dynamics simulations. Based on these results, and previous single-molecule data and cryo-EM reconstruction of the homologous φ29 motor, we propose an overall packaging model that is driven by helical-to-planar transitions of the ring motor. These transitions are coordinated by inter-subunit interactions that regulate catalytic and force-generating events. Stepwise ATP binding to individual subunits increase their affinity for the helical DNA phosphate backbone, resulting in distortion away from the planar ring towards a helical configuration, inducing mechanical strain. Subsequent sequential hydrolysis events alleviate the accumulated mechanical strain, allowing a stepwise return of the motor to the planar conformation, translocating DNA in the process. This type of helical-to-planar mechanism could serve as a general framework for ring ATPases.
- Dept. of Mechanical Engineering and Materials Science, Duke University, Durham, NC 27708, USA.
Organizational Affiliation: