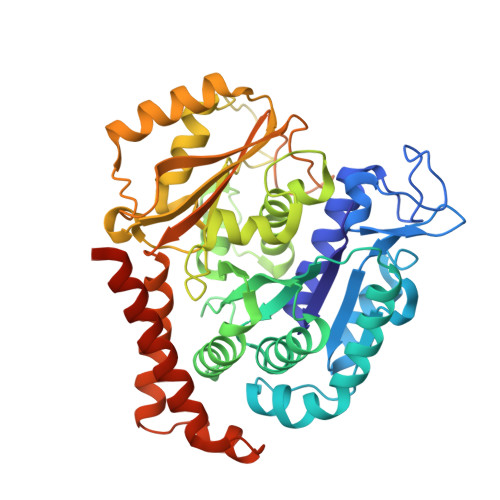
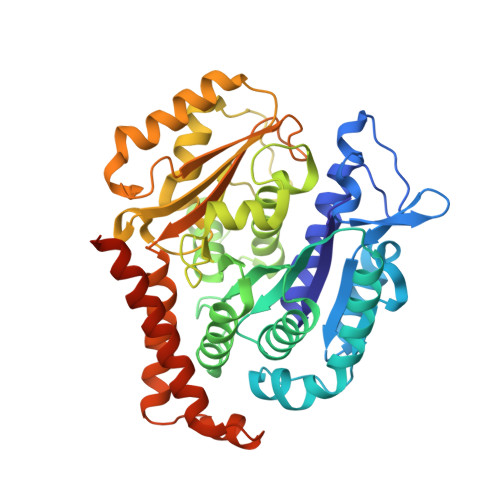
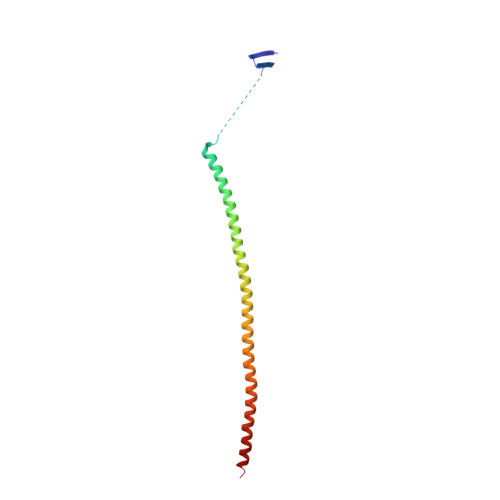
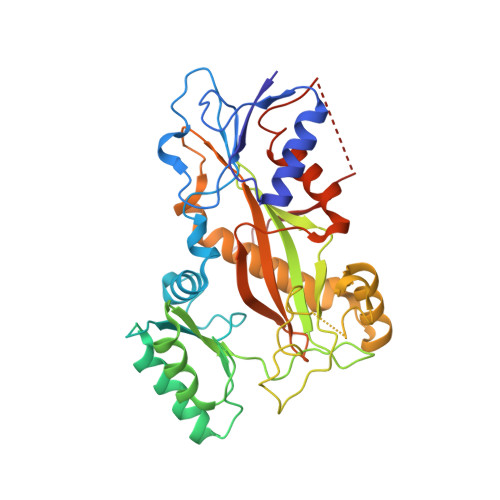
The X-ray structure of tubulysin analogue TGL in complex with tubulin and three possible routes for the development of next-generation tubulysin analogues.
Li, W., Tan, L., Zhang, Z., Xia, Q., Lei, D., Li, Y., Zhang, T., Zeng, S., Sima, X., Wang, Y.(2021) Biochem Biophys Res Commun 565: 29-35
- PubMed: 34090207 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbrc.2021.05.086
- Primary Citation Related Structures:
7EN3 - PubMed Abstract:
Microtubule-targeting agents (MTAs) are the most commonly used anti-cancer drugs. At least fourteen microtubule inhibitors and ten antibody drug conjugates (ADCs) linking MTAs are approved by FDA for clinical use in cancer therapy. In current research, we determined the crystal structure of tubulysin analogue TGL in complex with tubulin at a high resolution (2.65 Å). In addition, we summarized all of the previously published high-resolution crystal structures of ligands in the vinca site to provide structural insights for the rational design of the new vinca-site ligands. Moreover, based on the aligned results of the vinca site ligands, we provided three possible routes for designing new tubulysin analogues, namely macrocyclization between the N-14 side chain and the N-9 side chain, the hybird of tubulysin M and phomopsin A, and growing new aryl group at C-21. These designed structures will inspire the development of new MTAs or payloads in cancer therapy.
- State Key Laboratory of Biotherapy and Collaborative Innovation Center for Biotherapy, West China Hospital, Sichuan University, Chengdu, 610041, People's Republic of China. Electronic address: liwentingo@qq.com.
Organizational Affiliation: