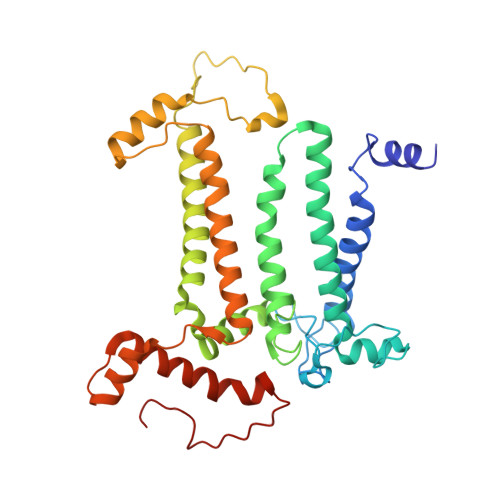
Structural insights into cyanobacterial photosystem II intermediates associated with Psb28 and Tsl0063.
Xiao, Y., Huang, G., You, X., Zhu, Q., Wang, W., Kuang, T., Han, G., Sui, S.F., Shen, J.R.(2021) Nat Plants 7: 1132-1142
- PubMed: 34226692 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41477-021-00961-7
- Primary Citation Related Structures:
7DXA, 7DXH - PubMed Abstract:
Photosystem II (PSII) is a multisubunit pigment-protein complex and catalyses light-induced water oxidation, leading to the conversion of light energy into chemical energy and the release of dioxygen. We analysed the structures of two Psb28-bound PSII intermediates, Psb28-RC47 and Psb28-PSII, purified from a psbV-deletion strain of the thermophilic cyanobacterium Thermosynechococcus vulcanus, using cryo-electron microscopy. Both Psb28-RC47 and Psb28-PSII bind one Psb28, one Tsl0063 and an unknown subunit. Psb28 is located at the cytoplasmic surface of PSII and interacts with D1, D2 and CP47, whereas Tsl0063 is a transmembrane subunit and binds at the side of CP47/PsbH. Substantial structural perturbations are observed at the acceptor side, which result in conformational changes of the quinone (Q B ) and non-haem iron binding sites and thus may protect PSII from photodamage during assembly. These results provide a solid structural basis for understanding the assembly process of native PSII.
- Photosynthesis Research Center, Key Laboratory of Photobiology, Institute of Botany, Chinese Academy of Sciences, Beijing, China.
Organizational Affiliation: