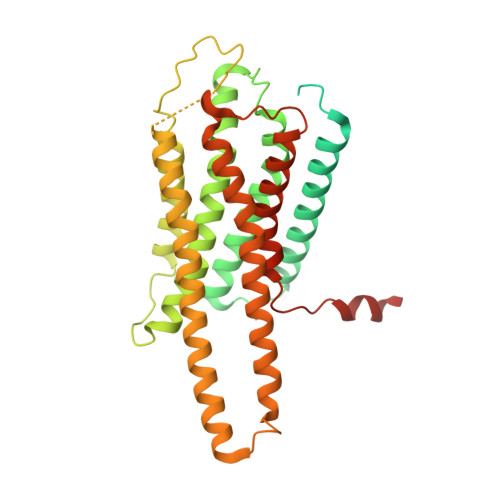
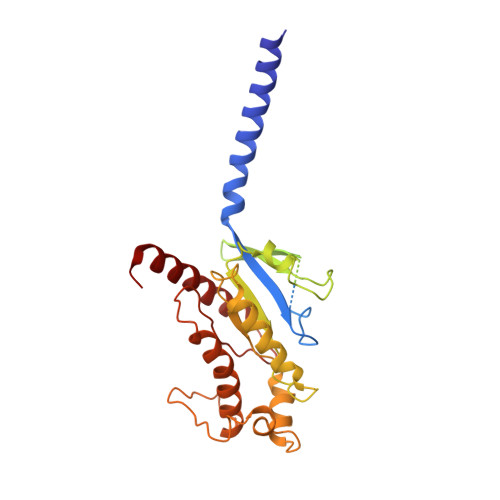
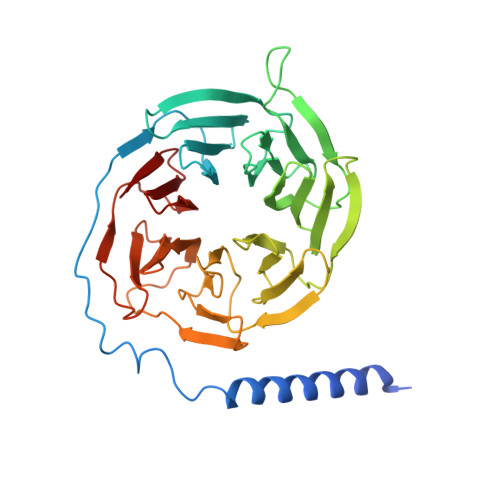
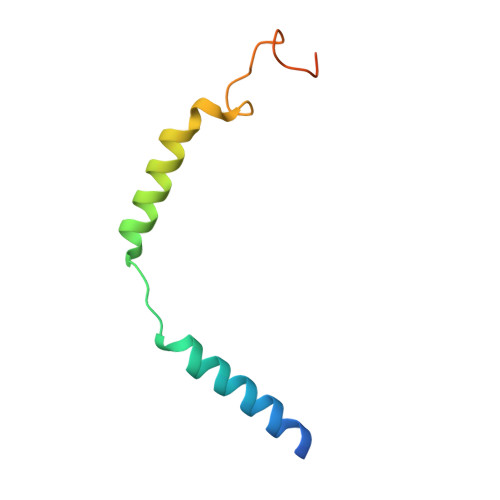
Cryo-EM structure of activated bile acids receptor TGR5 in complex with stimulatory G protein.
Chen, G., Wang, X., Ge, Y., Ma, L., Chen, Q., Liu, H., Du, Y., Ye, R.D., Hu, H., Ren, R.(2020) Signal Transduct Target Ther 5: 142-142
- PubMed: 32747649 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41392-020-00262-z
- Primary Citation Related Structures:
7BW0 - Kobilka Institute of Innovative Drug Discovery, School of Life and Health Sciences, The Chinese University of Hong Kong, 518172, Shenzhen, Guangdong, P.R. China.
Organizational Affiliation: