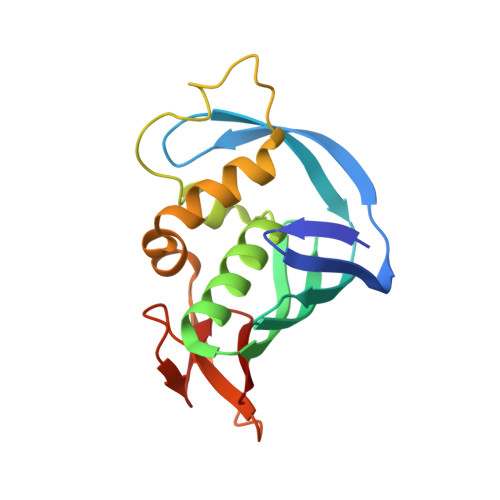
Structural and Thermodynamic Analysis of the Resistance Development to Pimodivir (VX-787), the Clinical Inhibitor of Cap Binding to PB2 Subunit of Influenza A Polymerase.
Gregor, J., Radilova, K., Brynda, J., Fanfrlik, J., Konvalinka, J., Kozisek, M.(2021) Molecules 26
- PubMed: 33673017 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.3390/molecules26041007
- Primary Citation Related Structures:
7AS0, 7AS1, 7AS2, 7AS3 - PubMed Abstract:
Influenza A virus (IAV) encodes a polymerase composed of three subunits: PA, with endonuclease activity, PB1 with polymerase activity and PB2 with host RNA five-prime cap binding site. Their cooperation and stepwise activation include a process called cap-snatching, which is a crucial step in the IAV life cycle. Reproduction of IAV can be blocked by disrupting the interaction between the PB2 domain and the five-prime cap. An inhibitor of this interaction called pimodivir (VX-787) recently entered the third phase of clinical trial; however, several mutations in PB2 that cause resistance to pimodivir were observed. First major mutation, F404Y, causing resistance was identified during preclinical testing, next the mutation M431I was identified in patients during the second phase of clinical trials. The mutation H357N was identified during testing of IAV strains at Centers for Disease Control and Prevention. We set out to provide a structural and thermodynamic analysis of the interactions between cap-binding domain of PB2 wild-type and PB2 variants bearing these mutations and pimodivir. Here we present four crystal structures of PB2-WT, PB2-F404Y, PB2-M431I and PB2-H357N in complex with pimodivir. We have thermodynamically analysed all PB2 variants and proposed the effect of these mutations on thermodynamic parameters of these interactions and pimodivir resistance development. These data will contribute to understanding the effect of these missense mutations to the resistance development and help to design next generation inhibitors.
- Institute of Organic Chemistry and Biochemistry of the Czech Academy of Sciences, Gilead Sciences and IOCB Research Center, Flemingovo n. 2, 16610 Prague 6, Czech Republic.
Organizational Affiliation: