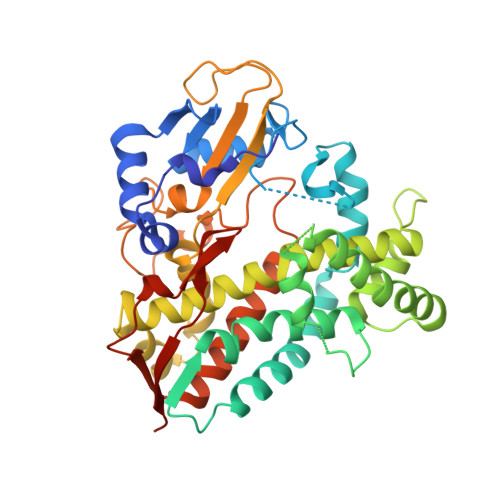
Structural comparison of the cytochrome P450 enzymes CYP106A1 and CYP106A2 provides insight into their differences in steroid conversion.
Carius, Y., Hutter, M., Kiss, F., Bernhardt, R., Lancaster, C.R.D.(2022) FEBS Lett 596: 3133-3144
- PubMed: 36151590 Search on PubMed
- DOI: https://doi.org/10.1002/1873-3468.14502
- Primary Citation Related Structures:
7Q9E, 7ZZL - PubMed Abstract:
Understanding the structural basis of the selectivity of steroid hydroxylation requires detailed structural and functional investigations on various steroid hydroxylases with different selectivities, such as the bacterial cytochrome P450 enzymes. Here, the crystal structure of the cytochrome P450 CYP106A1 from Priestia megaterium was solved. CYP106A1 exhibits a rare additional structural motif of a cytochrome P450, a sixth β-sheet. The protein was found in different unusual conformations corresponding to both open and closed forms even when crystallized without any known substrate. The structural comparison of CYP106A1 with the previously investigated CYP106A2, including docking studies for both isoforms with the substrate cortisol, reveals a completely different orientation of the steroid molecule in the active sites. This distinction convincingly explains the experimentally observed differences in substrate conversion and product formation by the two enzymes.
- Department of Structural Biology, Faculty of Medicine, Center of Human and Molecular Biology (ZHMB), Saarland University, Homburg, Germany.
Organizational Affiliation: