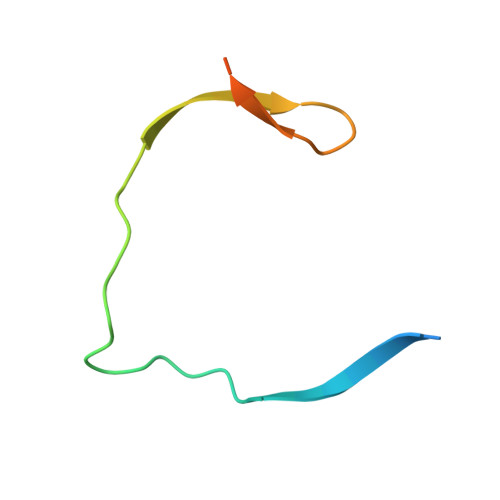
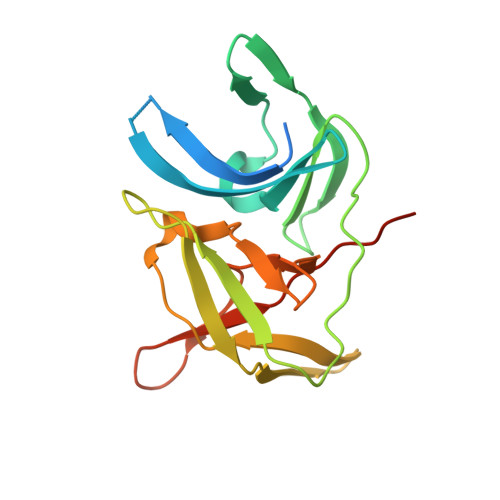
Synthesis and structural characterization of new macrocyclic inhibitors of the Zika virus NS2B-NS3 protease.
Huber, S., Braun, N.J., Schmacke, L.C., Murra, R., Bender, D., Hildt, E., Heine, A., Steinmetzer, T.(2024) Arch Pharm (Weinheim) : e2400250-e2400250
- PubMed: 38809037 Search on PubMed
- DOI: https://doi.org/10.1002/ardp.202400250
- Primary Citation Related Structures:
7ZPD, 7ZQ1, 7ZQF, 7ZTM, 7ZUM, 7ZV4, 7ZVV, 7ZW5, 7ZYS, 8A15, 8AQA, 8AQB, 8AQK - PubMed Abstract:
Three new series of macrocyclic active site-directed inhibitors of the Zika virus (ZIKV) NS2B-NS3 protease were synthesized. First, attempts were made to replace the basic P3 lysine residue of our previously described inhibitors with uncharged and more hydrophobic residues. This provided numerous compounds with inhibition constants between 30 and 50 nM. A stronger reduction of the inhibitory potency was observed when the P2 lysine was replaced by neutral residues, all of these inhibitors possess K i values >1 µM. However, it is possible to replace the P2 lysine with the less basic 3-aminomethylphenylalanine, which provides a similarly potent inhibitor of the ZIKV protease (K i = 2.69 nM). Crystal structure investigations showed that the P2 benzylamine structure forms comparable interactions with the protease as lysine. Twelve additional structures of these inhibitors in complex with the protease were determined, which explain many, but not all, SAR data obtained in this study. All individual modifications in the P2 or P3 position resulted in inhibitors with low antiviral efficacy in cell culture. Therefore, a third inhibitor series with combined modifications was synthesized; all of them contain a more hydrophobic d-cyclohexylalanine in the linker segment. At a concentration of 40 µM, two of these compounds possess similar antiviral potency as ribavirin at 100 µM. Due to their reliable crystallization in complex with the ZIKV protease, these cyclic compounds are very well suited for a rational structure-based development of improved inhibitors.
- Institute of Pharmaceutical Chemistry, Philipps University of Marburg, Marburg, Germany.
Organizational Affiliation: