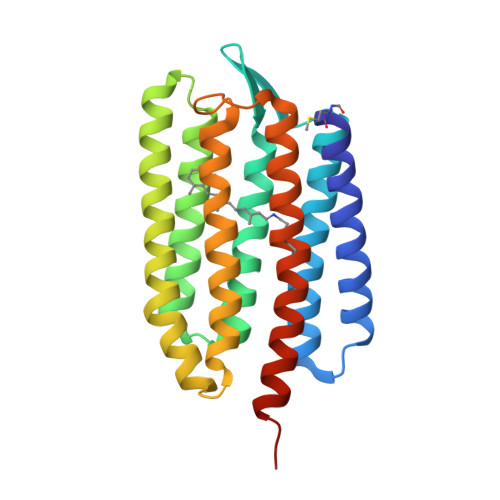
Structural insights into light-driven anion pumping in cyanobacteria.
Astashkin, R., Kovalev, K., Bukhdruker, S., Vaganova, S., Kuzmin, A., Alekseev, A., Balandin, T., Zabelskii, D., Gushchin, I., Royant, A., Volkov, D., Bourenkov, G., Koonin, E., Engelhard, M., Bamberg, E., Gordeliy, V.(2022) Nat Commun 13: 6460-6460
- PubMed: 36309497 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-022-34019-9
- Primary Citation Related Structures:
7ZOU, 7ZOV, 7ZOW, 7ZOY - PubMed Abstract:
Transmembrane ion transport is a key process in living cells. Active transport of ions is carried out by various ion transporters including microbial rhodopsins (MRs). MRs perform diverse functions such as active and passive ion transport, photo-sensing, and others. In particular, MRs can pump various monovalent ions like Na + , K + , Cl - , I - , NO 3 - . The only characterized MR proposed to pump sulfate in addition to halides belongs to the cyanobacterium Synechocystis sp. PCC 7509 and is named Synechocystis halorhodopsin (SyHR). The structural study of SyHR may help to understand what makes an MR pump divalent ions. Here we present the crystal structure of SyHR in the ground state, the structure of its sulfate-bound form as well as two photoreaction intermediates, the K and O states. These data reveal the molecular origin of the unique properties of the protein (exceptionally strong chloride binding and proposed pumping of divalent anions) and sheds light on the mechanism of anion release and uptake in cyanobacterial halorhodopsins. The unique properties of SyHR highlight its potential as an optogenetics tool and may help engineer different types of anion pumps with applications in optogenetics.
- Univ. Grenoble Alpes, CEA, CNRS, Institut de Biologie Structurale (IBS), Grenoble, France.
Organizational Affiliation: