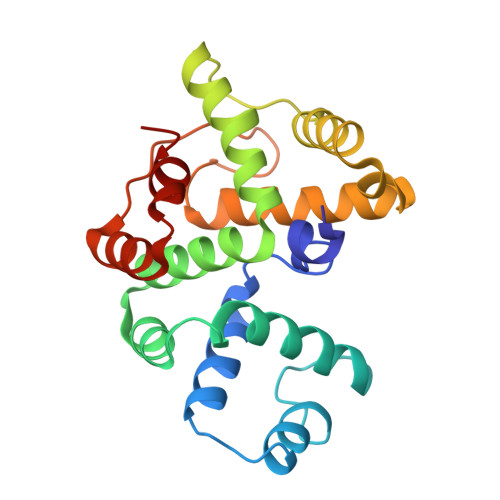
Crystal structure of the BREX phage defence protein BrxA.
Beck, I.N., Picton, D.M., Blower, T.R.(2022) Curr Res Struct Biol 4: 211-219
- PubMed: 35783086 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.crstbi.2022.06.001
- Primary Citation Related Structures:
7ZGE - PubMed Abstract:
Bacteria are constantly challenged by bacteriophage (phage) infection and have developed multitudinous and varied resistance mechanisms. Bacteriophage Exclusion (BREX) systems protect from phage infection by generating methylation patterns at non-palindromic 6 bp sites in host bacterial DNA, to distinguish and block replication of non-self DNA. Type 1 BREX systems are comprised of six conserved core genes. Here, we present the first reported structure of a BREX core protein, BrxA from the phage defence island of Escherichia fergusonii ATCC 35469 plasmid pEFER, solved to 2.09 Å. BrxA is a monomeric protein in solution, with an all α-helical globular fold. Conservation of surface charges and structural homology modelling against known phage defence systems highlighted that BrxA contains two helix-turn-helix motifs, juxtaposed by 180°, positioned to bind opposite sides of a DNA major groove. BrxA was subsequently shown to bind dsDNA. This new understanding of BrxA structure, and first indication of BrxA biological activity, suggests a conserved mode of DNA-recognition has become widespread and implemented by diverse phage defence systems.
- Department of Biosciences, Durham University, Stockton Road, Durham, DH1 3LE, UK.
Organizational Affiliation: