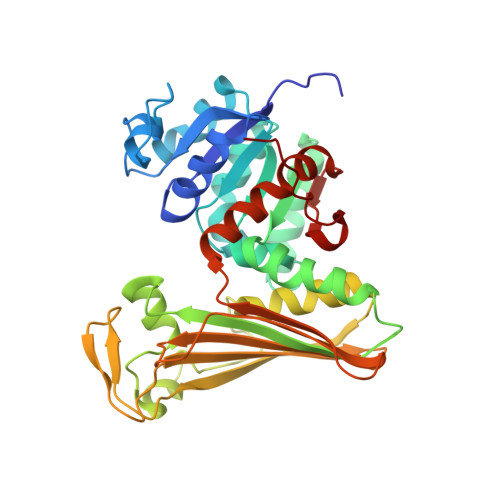
Structure and Mutation of the Native Amine Dehydrogenase MATOUAmDH2.
Bennett, M., Ducrot, L., Vergne-Vaxelaire, C., Grogan, G.(2022) Chembiochem 23: e202200136-e202200136
- PubMed: 35349204 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/cbic.202200136
- Primary Citation Related Structures:
7R09, 7ZBO - PubMed Abstract:
Native amine dehydrogenases (nat-AmDHs) have recently emerged as a potentially valuable new reservoir of enzymes for the sustainable and selective synthesis of chiral amines, catalyzing the NAD(P)H-dependent ammoniation of carbonyl compounds with high activity and selectivity. MATOUAmDH2, recently identified from the Marine Atlas of Tara Oceans Unigenes (MATOUv1) database of eukaryotic genes, displays exceptional catalytic performance against its best identified substrate, isobutyraldehyde, as well as having broader substrate scope than other nat-AmDHs. In the interests of providing a platform for the rational engineering of this and other nat-AmDHs, we have determined the structure of MATOUAmDH2 in complex with NADP + and also with the cofactor and cyclohexylamine. Monomers within the structure are representative of more open and closed conformations of the enzyme and illustrate the profound changes undergone by nat-AmDHs during the catalytic cycle. An alanine screen of active site residues revealed that M215A and L180A are more active than the wild-type enzyme for the amination of cyclohexanone with ammonia and methylamine respectively; the latter suggests that AmDHs have the potential to be engineered for the improved production of secondary amines.
- Department of Chemistry, University of York, Heslington, York YO10 5DD, UK.
Organizational Affiliation: