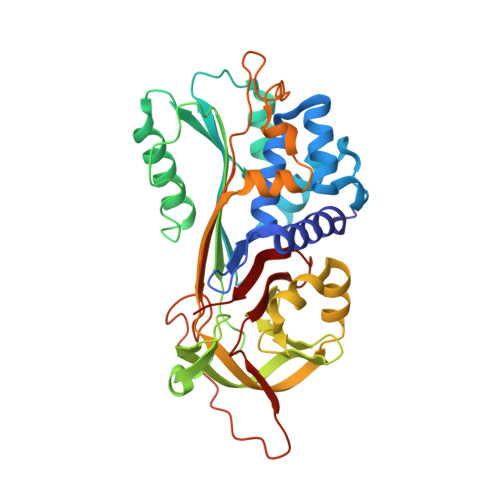
Conformational transition of the Ixodes ricinus salivary serpin Iripin-4.
Kascakova, B., Kotal, J., Havlickova, P., Vopatkova, V., Prudnikova, T., Grinkevich, P., Kuty, M., Chmelar, J., Kuta Smatanova, I.(2023) Acta Crystallogr D Struct Biol 79: 409-419
- PubMed: 37092969 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2059798323002322
- Primary Citation Related Structures:
7ZAS, 7ZBF - PubMed Abstract:
Iripin-4, one of the many salivary serpins from Ixodes ricinus ticks with an as-yet unexplained function, crystallized in two different structural conformations, namely the native partially relaxed state and the cleaved serpin. The native structure was solved at a resolution of 2.3 Å and the structure of the cleaved conformation was solved at 2.0 Å resolution. Furthermore, structural changes were observed when the reactive-centre loop transitioned from the native conformation to the cleaved conformation. In addition to this finding, it was confirmed that Glu341 represents a primary substrate-recognition site for the inhibitory mechanism. The presence of glutamate instead of the typical arginine in the P1 recognition site of all structurally characterized I. ricinus serpins (PDB entries 7b2t, 7pmu and 7ahp), except for the tyrosine in the P1 site of Iripin-2 (formerly IRS-2; PDB entry 3nda), would explain the absence of inhibition of the tested proteases that cleave their substrate after arginine. Further research on Iripin-4 should focus on functional analysis of this interesting serpin.
- Department of Chemistry, University of South Bohemia in České Budějovice, 370 05 České Budějovice, Czech Republic.
Organizational Affiliation: