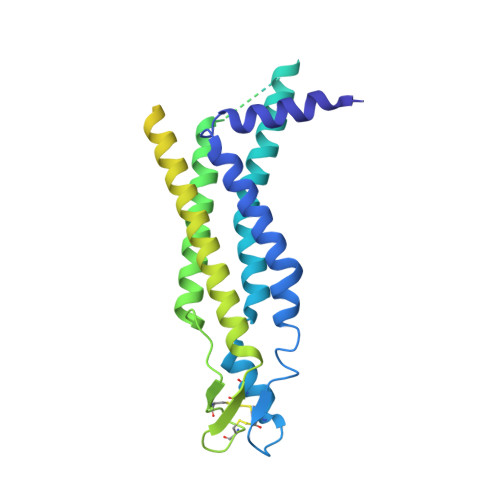
Structure of the connexin-43 gap junction channel in a putative closed state.
Qi, C., Acosta Gutierrez, S., Lavriha, P., Othman, A., Lopez-Pigozzi, D., Bayraktar, E., Schuster, D., Picotti, P., Zamboni, N., Bortolozzi, M., Gervasio, F.L., Korkhov, V.M.(2023) Elife 12
- PubMed: 37535063 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.7554/eLife.87616
- Primary Citation Related Structures:
7Z1T, 7Z22, 7Z23 - PubMed Abstract:
Gap junction channels (GJCs) mediate intercellular communication by connecting two neighbouring cells and enabling direct exchange of ions and small molecules. Cell coupling via connexin-43 (Cx43) GJCs is important in a wide range of cellular processes in health and disease (Churko and Laird, 2013; Liang et al., 2020; Poelzing and Rosenbaum, 2004), yet the structural basis of Cx43 function and regulation has not been determined until now. Here, we describe the structure of a human Cx43 GJC solved by cryo-EM and single particle analysis at 2.26 Å resolution. The pore region of Cx43 GJC features several lipid-like densities per Cx43 monomer, located close to a putative lateral access site at the monomer boundary. We found a previously undescribed conformation on the cytosolic side of the pore, formed by the N-terminal domain and the transmembrane helix 2 of Cx43 and stabilized by a small molecule. Structures of the Cx43 GJC and hemichannels (HCs) in nanodiscs reveal a similar gate arrangement. The features of the Cx43 GJC and HC cryo-EM maps and the channel properties revealed by molecular dynamics simulations suggest that the captured states of Cx43 are consistent with a closed state.
- Institute of Molecular Biology and Biophysics, ETH Zurich, Zurich, Switzerland.
Organizational Affiliation: