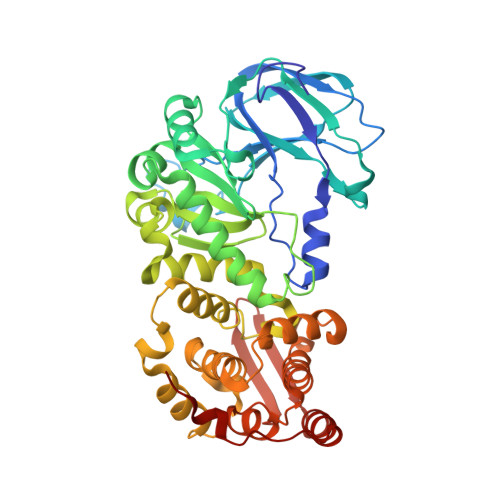
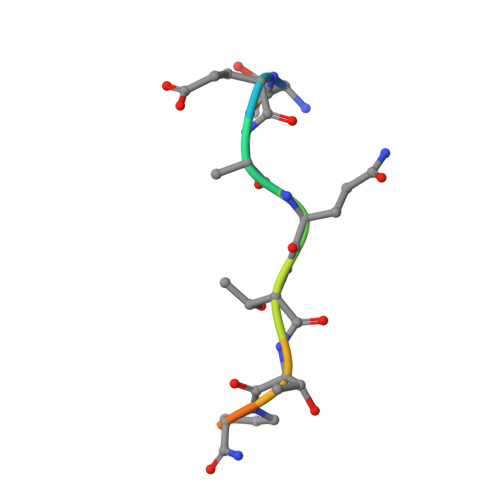
Structural and mechanistic insights into the cleavage of clustered O-glycan patches-containing glycoproteins by mucinases of the human gut.
Taleb, V., Liao, Q., Narimatsu, Y., Garcia-Garcia, A., Companon, I., Borges, R.J., Gonzalez-Ramirez, A.M., Corzana, F., Clausen, H., Rovira, C., Hurtado-Guerrero, R.(2022) Nat Commun 13: 4324-4324
- PubMed: 35882872 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-022-32021-9
- Primary Citation Related Structures:
7YX8 - PubMed Abstract:
Mucinases of human gut bacteria cleave peptide bonds in mucins strictly depending on the presence of neighboring O-glycans. The Akkermansia muciniphila AM0627 mucinase cleaves specifically in between contiguous (bis) O-glycans of defined truncated structures, suggesting that this enzyme may recognize clustered O-glycan patches. Here, we report the structure and molecular mechanism of AM0627 in complex with a glycopeptide containing a bis-T (Galβ1-3GalNAcα1-O-Ser/Thr) O-glycan, revealing that AM0627 recognizes both the sugar moieties and the peptide sequence. AM0627 exhibits preference for bis-T over bis-Tn (GalNAcα1-O-Ser/Thr) O-glycopeptide substrates, with the first GalNAc residue being essential for cleavage. AM0627 follows a mechanism relying on a nucleophilic water molecule and a catalytic base Glu residue. Structural comparison among mucinases identifies a conserved Tyr engaged in sugar-π interactions in both AM0627 and the Bacteroides thetaiotaomicron BT4244 mucinase as responsible for the common activity of these two mucinases with bis-T/Tn substrates. Our work illustrates how mucinases through tremendous flexibility adapt to the diversity in distribution and patterns of O-glycans on mucins.
- Institute of Biocomputation and Physics of Complex Systems, University of Zaragoza, Mariano Esquillor s/n, Campus Rio Ebro, Edificio I+D, Zaragoza, Spain.
Organizational Affiliation: