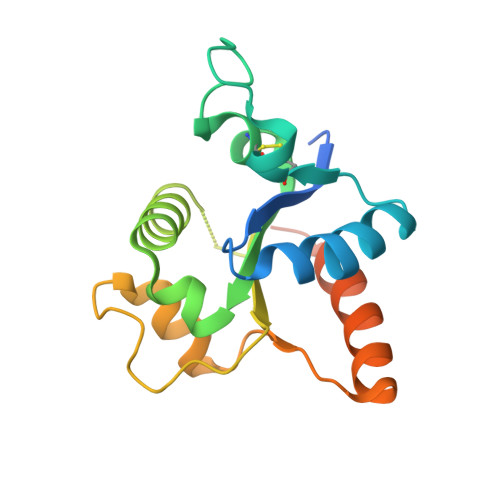
Structural analysis of the Toll-like receptor 15 TIR domain.
Ko, K.Y., Song, W.S., Park, J., Lee, G.S., Yoon, S.I.(2023) IUCrJ 10: 352-362
- PubMed: 37079400 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2052252523002956
- Primary Citation Related Structures:
7YLF, 7YLG - PubMed Abstract:
Toll-like receptors (TLRs) activate innate immunity in response to pathogen-associated molecular patterns (PAMPs). The ectodomain of a TLR directly senses a PAMP and the intracellular TIR domain dimerizes to initiate a signaling cascade. The TIR domains of TLR6 and TLR10, which belong to the TLR1 subfamily, have been structurally characterized in a dimer, whereas those of other subfamilies, including TLR15, have not been explored at the structural or molecular level. TLR15 is a TLR unique to birds and reptiles that responds to virulence-associated fungal and bacterial proteases. To reveal how the TLR15 TIR domain (TLR15 TIR ) triggers signaling, the crystal structure of TLR15 TIR was determined in a dimeric form and a mutational study was performed. TLR15 TIR forms a one-domain structure in which a five-stranded β-sheet is decorated by α-helices, as shown for TLR1 subfamily members. TLR15 TIR exhibits substantial structural differences from other TLRs at the BB and DD loops and αC2 helix that are involved in dimerization. As a result, TLR15 TIR is likely to form a dimeric structure that is unique in its intersubunit orientation and the contribution of each dimerizing region. Further comparative analysis of TIR structures and sequences provides insights into the recruitment of a signaling adaptor protein by TLR15 TIR .
- Division of Biomedical Convergence, Kangwon National University, Chuncheon, 24341, Republic of Korea.
Organizational Affiliation: