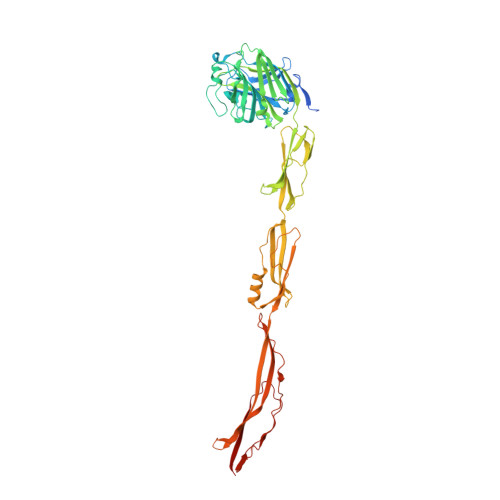Cell-surface protein YwfG of Lactococcus lactis binds to alpha-1,2-linked mannose.
Tsuchiya, W., Fujimoto, Z., Inagaki, N., Nakagawa, H., Tanaka, M., Kimoto-Nira, H., Yamazaki, T., Suzuki, C.(2023) PLoS One 18: e0273955-e0273955
- PubMed: 36602978 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pone.0273955
- Primary Citation Related Structures:
7YL4, 7YL5, 7YL6 - PubMed Abstract:
Lactococcus lactis strains are used as starter cultures in the production of fermented dairy and vegetable foods, but the species also occurs in other niches such as plant material. Lactococcus lactis subsp. lactis G50 (G50) is a plant-derived strain and potential candidate probiotics. Western blotting of cell-wall proteins using antibodies generated against whole G50 cells detected a 120-kDa protein. MALDI-TOF MS analysis identified it as YwfG, a Leu-Pro-any-Thr-Gly cell-wall-anchor-domain-containing protein. Based on a predicted domain structure, a recombinant YwfG variant covering the N-terminal half (aa 28-511) of YwfG (YwfG28-511) was crystallized and the crystal structure was determined. The structure consisted of an L-type lectin domain, a mucin-binding protein domain, and a mucus-binding protein repeat. Recombinant YwfG variants containing combinations of these domains (YwfG28-270, YwfG28-336, YwfG28-511, MubR4) were prepared and their interactions with monosaccharides were examined by isothermal titration calorimetry; the only interaction observed was between YwfG28-270, which contained the L-type lectin domain, and d-mannose. Among four mannobioses, α-1,2-mannobiose had the highest affinity for YwfG28-270 (dissociation constant = 34 μM). YwfG28-270 also interacted with yeast mannoproteins and yeast mannan. Soaking of the crystals of YwfG28-511 with mannose or α-1,2-mannobiose revealed that both sugars bound to the L-type lectin domain in a similar manner, although the presence of the mucin-binding protein domain and the mucus-binding protein repeat within the recombinant protein inhibited the interaction between the L-type lectin domain and mannose residues. Three of the YwfG variants (except MubR4) induced aggregation of yeast cells. Strain G50 also induced aggregation of yeast cells, which was abolished by deletion of ywfG from G50, suggesting that surface YwfG contributes to the interaction with yeast cells. These findings provide new structural and functional insights into the interaction between L. lactis and its ecological niche via binding of the cell-surface protein YwfG with mannose.
- Research Center for Advanced Analysis, National Agriculture and Food Research Organization (NARO), Tsukuba, Ibaraki, Japan.
Organizational Affiliation: