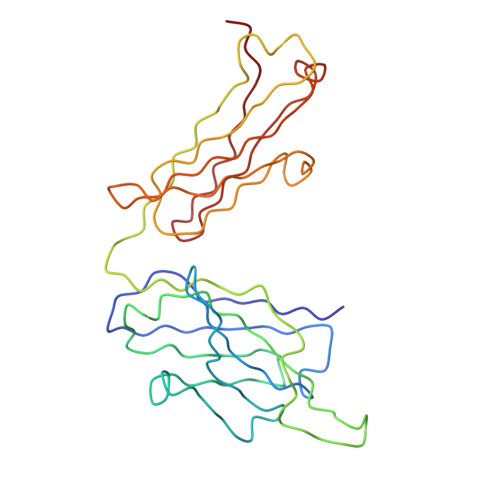
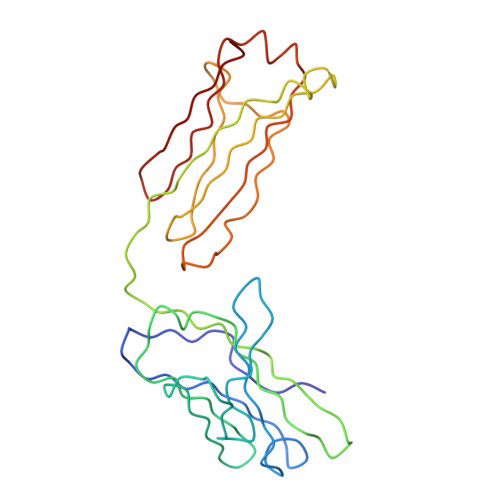
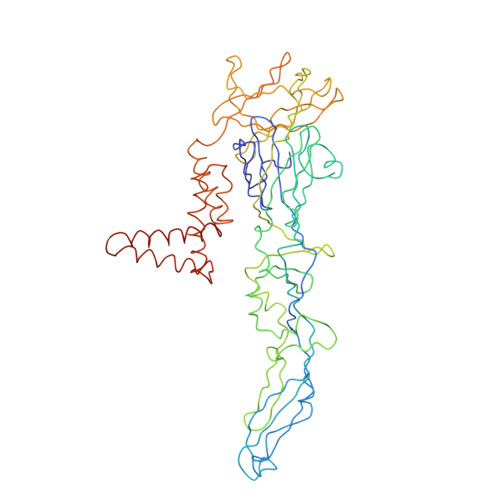
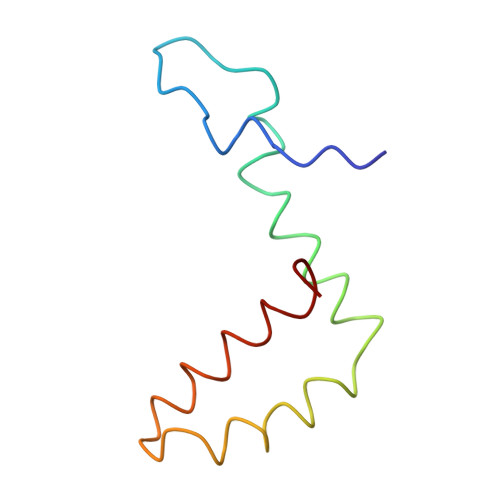
Structure and neutralization mechanism of a human antibody targeting a complex Epitope on Zika virus.
Adams, C., Carbaugh, D.L., Shu, B., Ng, T.S., Castillo, I.N., Bhowmik, R., Segovia-Chumbez, B., Puhl, A.C., Graham, S., Diehl, S.A., Lazear, H.M., Lok, S.M., de Silva, A.M., Premkumar, L.(2023) PLoS Pathog 19: e1010814-e1010814
- PubMed: 36626401 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.ppat.1010814
- Primary Citation Related Structures:
7YAR, 8DV6 - PubMed Abstract:
We currently have an incomplete understanding of why only a fraction of human antibodies that bind to flaviviruses block infection of cells. Here we define the footprint of a strongly neutralizing human monoclonal antibody (mAb G9E) with Zika virus (ZIKV) by both X-ray crystallography and cryo-electron microscopy. Flavivirus envelope (E) glycoproteins are present as homodimers on the virion surface, and G9E bound to a quaternary structure epitope spanning both E protomers forming a homodimer. As G9E mainly neutralized ZIKV by blocking a step after viral attachment to cells, we tested if the neutralization mechanism of G9E was dependent on the mAb cross-linking E molecules and blocking low-pH triggered conformational changes required for viral membrane fusion. We introduced targeted mutations to the G9E paratope to create recombinant antibodies that bound to the ZIKV envelope without cross-linking E protomers. The G9E paratope mutants that bound to a restricted epitope on one protomer poorly neutralized ZIKV compared to the wild-type mAb, demonstrating that the neutralization mechanism depended on the ability of G9E to cross-link E proteins. In cell-free low pH triggered viral fusion assay, both wild-type G9E, and epitope restricted paratope mutant G9E bound to ZIKV but only the wild-type G9E blocked fusion. We propose that, beyond antibody binding strength, the ability of human antibodies to cross-link E-proteins is a critical determinant of flavivirus neutralization potency.
- Department of Microbiology and Immunology, University of North Carolina School of Medicine, Chapel Hill, North Carolina, United States of America.
Organizational Affiliation: