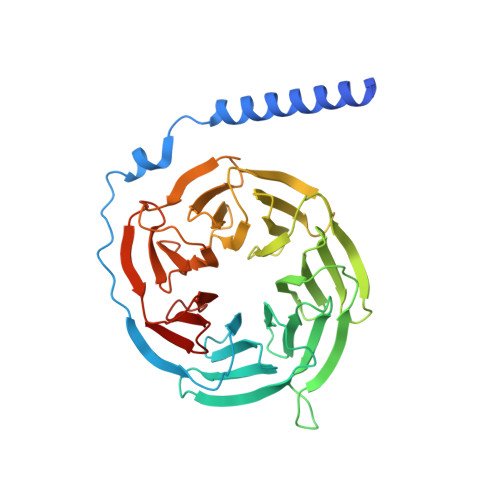
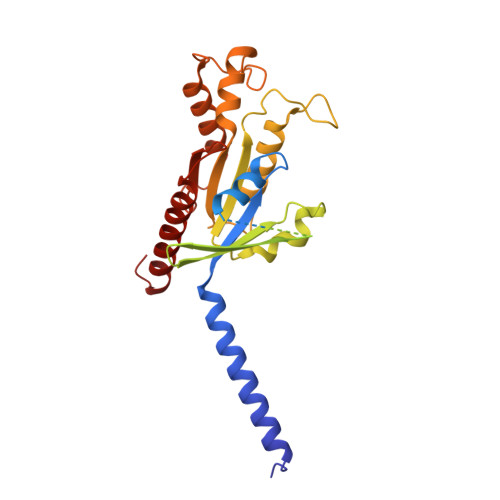
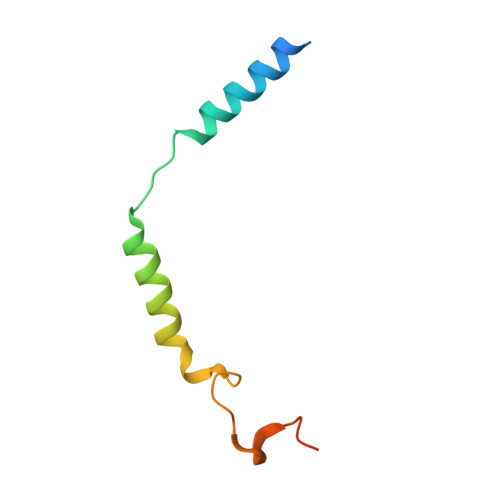
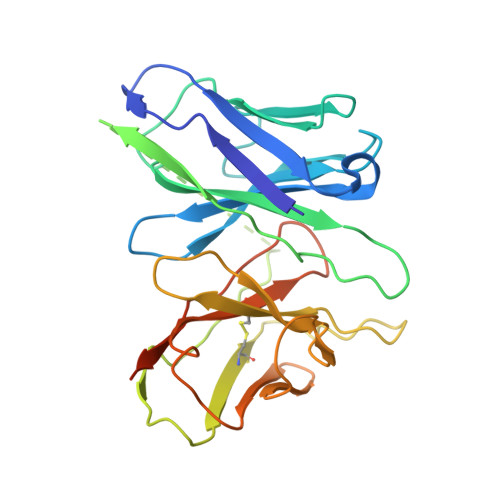
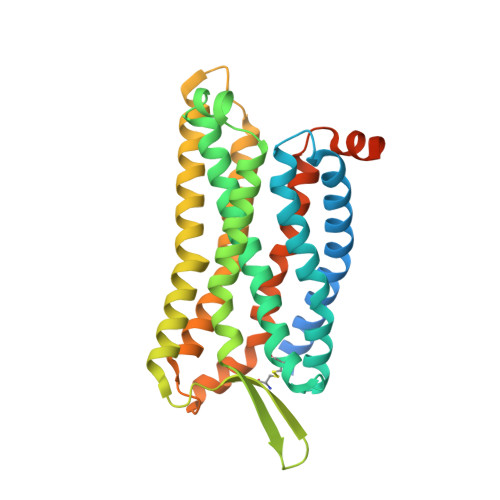
Mechanism of activation and biased signaling in complement receptor C5aR1.
Feng, Y., Zhao, C., Deng, Y., Wang, H., Ma, L., Liu, S., Tian, X., Wang, B., Bin, Y., Chen, P., Yan, W., Fu, P., Shao, Z.(2023) Cell Res 33: 312-324
- PubMed: 36806352 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41422-023-00779-2
- Primary Citation Related Structures:
7Y64, 7Y65, 7Y66, 7Y67 - PubMed Abstract:
The complement system plays an important role in the innate immune response to invading pathogens. The complement fragment C5a is one of its important effector components and exerts diverse physiological functions through activation of the C5a receptor 1 (C5aR1) and associated downstream G protein and β-arrestin signaling pathways. Dysfunction of the C5a-C5aR1 axis is linked to numerous inflammatory and immune-mediated diseases, but the structural basis for activation and biased signaling of C5aR1 remains elusive. Here, we present cryo-electron microscopy structures of the activated wild-type C5aR1-G i protein complex bound to each of the following: C5a, the hexapeptidic agonist C5a pep , and the G protein-biased agonist BM213. The structures reveal the landscape of the C5a-C5aR1 interaction as well as a common motif for the recognition of diverse orthosteric ligands. Moreover, combined with mutagenesis studies and cell-based pharmacological assays, we deciphered a framework for biased signaling using different peptide analogs and provided insight into the activation mechanism of C5aR1 by solving the structure of C5aR1 I116A mutant-G i signaling activation complex induced by C089, which exerts antagonism on wild-type C5aR1. In addition, unusual conformational changes in the intracellular end of transmembrane domain 7 and helix 8 upon agonist binding suggest a differential signal transduction process. Collectively, our study provides mechanistic understanding into the ligand recognition, biased signaling modulation, activation, and G i protein coupling of C5aR1, which may facilitate the future design of therapeutic agents.
- Division of Nephrology and Kidney Research Institute, State Key Laboratory of Biotherapy and Cancer Center, West China Hospital, Sichuan University, Chengdu, Sichuan, China.
Organizational Affiliation: